MEGATRON: Meta-Learning for Next-Generation Advanced Technology Realization & Acceleration
Keywords: Meta-learning, Data Reuse, Few-shot Learning, Electrical Impedance Tomography, Breast Cancer Detection
Introduction
Electrical Impedance Tomography (EIT) has emerged as a promising non-invasive imaging modality that offers significant advantages over conventional screening methods. EIT technology provides portable, radiation-free imaging capabilities that can potentially transform point-of-care breast cancer screening, particularly in resource-limited settings or for patients who experience discomfort with traditional mammography procedures (Halter et al., 2008). The fundamental principle underlying EIT involves the measurement of electrical impedance variations within biological tissues, which can reveal pathological changes indicative of malignant processes. Recent developments in EIT technology have demonstrated considerable potential for early breast cancer detection, with several studies showing promising results when combined with advanced signal processing and machine learning algorithms (Polydorides & Lionheart, 2002; Lionheart, 2004). However, the clinical translation and widespread adoption of EIT-based diagnostic tools face a critical bottleneck: the scarcity of sufficient real-world EIT datasets required for training robust machine learning models capable of accurate tumor detection and classification.
The development of effective machine learning algorithms for medical imaging typically requires extensive datasets comprising thousands to millions of labeled examples to achieve clinically acceptable performance levels. Traditional approaches to addressing this requirement in emerging imaging modalities like EIT have relied heavily on synthetic data generation and physics-based simulations (Liu et al., 2018). While synthetic data has proven useful for initial algorithm development, it inherently fails to capture the pathological variability and demographic diversity present in real-world clinical environments, resulting in models that may perform well in idealized laboratory settings but demonstrate limited generalizability to actual patient populations. The challenges associated with synthetic data generation extend beyond mere representational limitations. Creating realistic synthetic EIT datasets requires extensive computational resources, deep understanding of the underlying physics, and careful consideration of noise characteristics that mirror real-world acquisition conditions. Concurrently, vast amounts of conventional medical imaging data from established modalities such as mammography, ultrasound, MRI, and X-ray remain underutilized in the development of novel imaging technologies. These datasets, accumulated over decades of clinical practice and research, represent a rich repository of expertly annotated pathological and normal cases that could potentially accelerate the development of emerging diagnostic modalities if appropriately leveraged.
Meta-learning, often described as "learning to learn," presents a compelling solution to the data scarcity challenge facing novel medical imaging technologies (Finn et al., 2017). This paradigm enables machine learning models to rapidly adapt to new tasks with minimal training data by leveraging knowledge gained from related tasks during a meta-training phase. In the context of medical imaging, meta-learning offers the possibility of training models on abundant conventional datasets and subsequently fine-tuning them for novel modalities with limited available data. The application of meta-learning to medical imaging represents a paradigm shift from traditional single-task learning approaches toward more generalizable, adaptable systems that can extract universal feature representations applicable across different imaging modalities. This approach is particularly relevant for emerging technologies like EIT, where the fundamental task of identifying pathological tissue changes shares commonalities with established imaging modalities, despite differences in acquisition methods and image characteristics.
This study presents MEGATRON: Meta-Learning for Next-Generation Advanced Technology Realization & Acceleration, a comprehensive framework designed to address the data scarcity limitations hindering the clinical deployment of EIT-based breast cancer detection systems. The primary objective of this research is to demonstrate the feasibility and effectiveness of using meta-learning approaches trained on conventional medical imaging datasets to enable accurate few-shot tumor detection on limited EIT data. We present an automated, robust ETL (Extract, Transform, Load) pipeline capable of ingesting and standardizing heterogeneous medical imaging data from multiple open-source repositories, formats, and modalities, while introducing a novel meta-learning framework specifically designed for medical imaging tasks that can effectively transfer knowledge from conventional imaging modalities to emerging technologies like EIT. Our study provides empirical evidence for the effectiveness of cross-modal knowledge transfer between different medical imaging modalities, demonstrating that features learned from conventional imaging can enhance performance on novel imaging tasks. Additionally, we contribute a validated, reproducible open-source framework that can be applied beyond EIT to accelerate the development of various emerging diagnostic technologies facing similar data scarcity challenges, establishing a practical pathway for translating research-stage imaging technologies into clinically deployable tools by leveraging existing data resources.
The implications of this research extend beyond breast cancer detection and EIT technology. The meta-learning framework presented here addresses a fundamental challenge in medical technology development, that is, the need to validate and deploy novel diagnostic tools rapidly while ensuring clinical accuracy and reliability. By demonstrating effective knowledge transfer across imaging modalities, this work opens new avenues for accelerating medical innovation and improving patient outcomes through faster technology translation. The remainder of this paper presents our methodology, experimental results, and analysis of the MEGATRON framework's performance in enabling few-shot learning for EIT-based breast cancer detection, along with a discussion of broader applications and future research directions.
Methodology
Our methodology is organized into two interconnected pipelines.
Data Processing Pipeline:
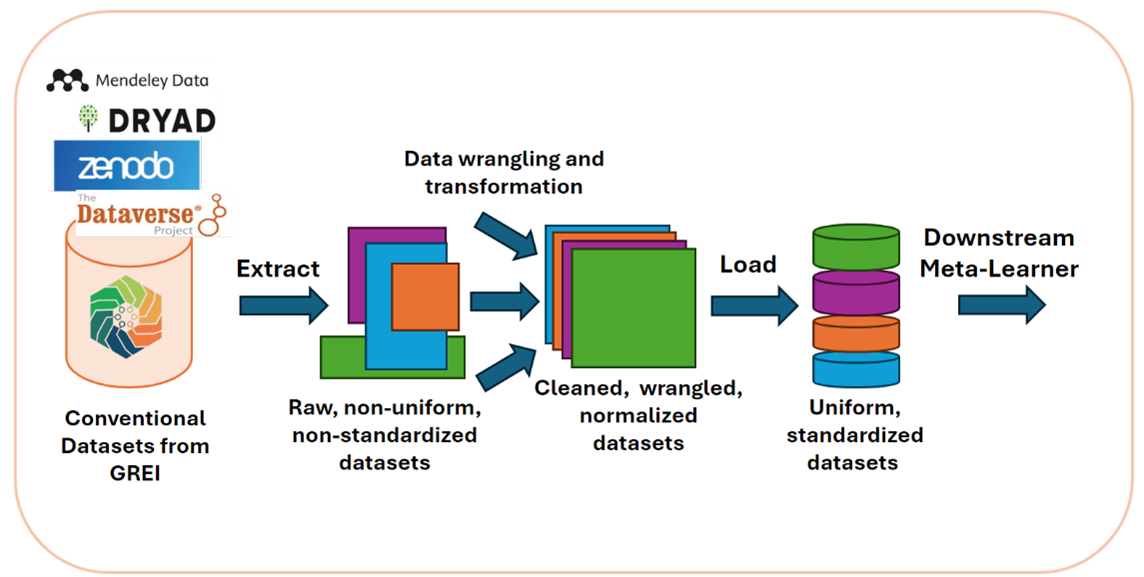
To enable robust training across diverse medical imaging modalities, we design a scalable data pipeline that integrates different repositories from GREI e.g., Dataverse, Figshare, Mendeley, Zenodo. The pipeline complies with FAIR principles, enabling automated ingestion, transformation, and storage of large, heterogeneous medical imaging datasets. The steps are:
-
Extract – Ingest raw imaging datasets from conventional modalities (e.g., MRI, X-ray, ultrasound) and Electrical Impedance Tomography (EIT) from remote URLs and online repositories.
-
Transform – Normalize metadata formats, ensure consistent labeling, and remove corrupted or incomplete samples. Apply domain-specific transformations (e.g., resizing, intensity normalization, coordinate transforms) to unify datasets for downstream training.
-
Load - Store the processed data in a standardized directory structure to enable ease of loading using standard dataloaders.
This pipeline ensures that the downstream model is trained on high-quality, standardized input that can generalize across modalities.
Training Pipeline:
The training strategy follows a staged approach that combines meta-training, meta-validation and fine-tuning for task-specific adaptation:
-
Meta-training – Train a base learner on large-scale conventional medical imaging datasets using an episodic training schedule to mimic few-shot learning scenarios.
-
Meta-Validation - Evaluate the meta-model’s performance on validation sets for each task using standard performance metrics for detection tasks e.g., mean Average Precision at 50% IoU, Recall, Precision, F1-score.
-
Fine-tuning – Adapt the meta-trained model to the target EIT dataset, enabling domain-specific optimization while retaining generalization ability.
The training pipeline ensures easy and scalable integration with processed data, enabling meta-learning from a variety of well-established medical imaging datasets and fine-tuning on data from low-resource domains like EIT. The pipeline periodically stores model checkpoints and optimizer states and records model performance on validation sets.
Requirements
Table 1: Hardware and software minimum requirements.
| Item | Minimum Required |
|---|---|
| GPU | CUDA-enabled with compute capability >=3.0 |
| CUDA Toolkit | 11.8 |
| Storage | 1 TB |
| vRAM | 40 GB |
| Linux Distro | Ubuntu 22.04 |
| Conda | 25.1.1 |
| Python | 3.10 |
Datasets
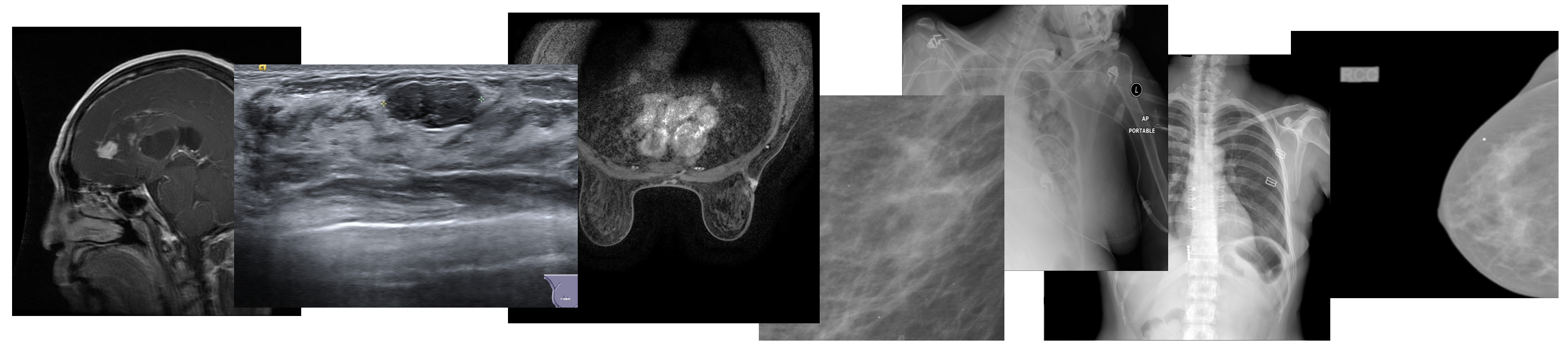

Table 1: A summary of the 8 datasets used to train a general-purpose meta-model. Datasets 1-7 are using during meta-model training, while dataset 8 is used to fine-tune the model and evaluate its few-shot object detection capabilities.
| Name | Pathology | Annotation Type | Classes | Image Format | No. of Samples | Source | DOI |
|---|---|---|---|---|---|---|---|
| Deepsight-2d-Mammogram | Breast Cancer | Detection | 3 | npz | 161589 | Dataverse | doi:10.7910/DVN/KXJCIU |
| MRI-Brain-Tumor | Brain Cancer | Detection | 4 | mat, jpg | 10087 | Figshare, Mendeley | doi:10.6084/m9.figshare.1512427, doi:10.17632/zp67tkpj2y.1 |
| Breast-Ultrasound | Breast Cancer | Segmentation | 3 | png | 683 | Mendeley | doi:10.17632/7fvgj4jsp7.3 |
| CBIS-DDSM-Mammogram | Breast Cancer | Segmentation | 2 | jpg, png | 3568 | Zenodo | doi:10.7937/K9/TCIA.2016.7O02S9CY |
| Advanced-MRI-Breast-Lesions | Breast Cancer | Segmentation | 2 | dcm | 632 | TCIA | doi:10.7937/C7X1-YN57 |
| Chest-xray | Pneumonia | Detection | 14 | png | 18000 | Zenodo | doi:10.5281/zenodo.12721389 |
| RSNA-pneumonia | Pneumonia | Detection | 2 | dcm | 29684 | RSNA | doi:10.1148/ryai.2019180041 |
| EIT-Novel-Data (Ours) | Breast Cancer | Detection | 2 | png | 40 | Zenodo | doi:10.5281/zenodo.17001019 |
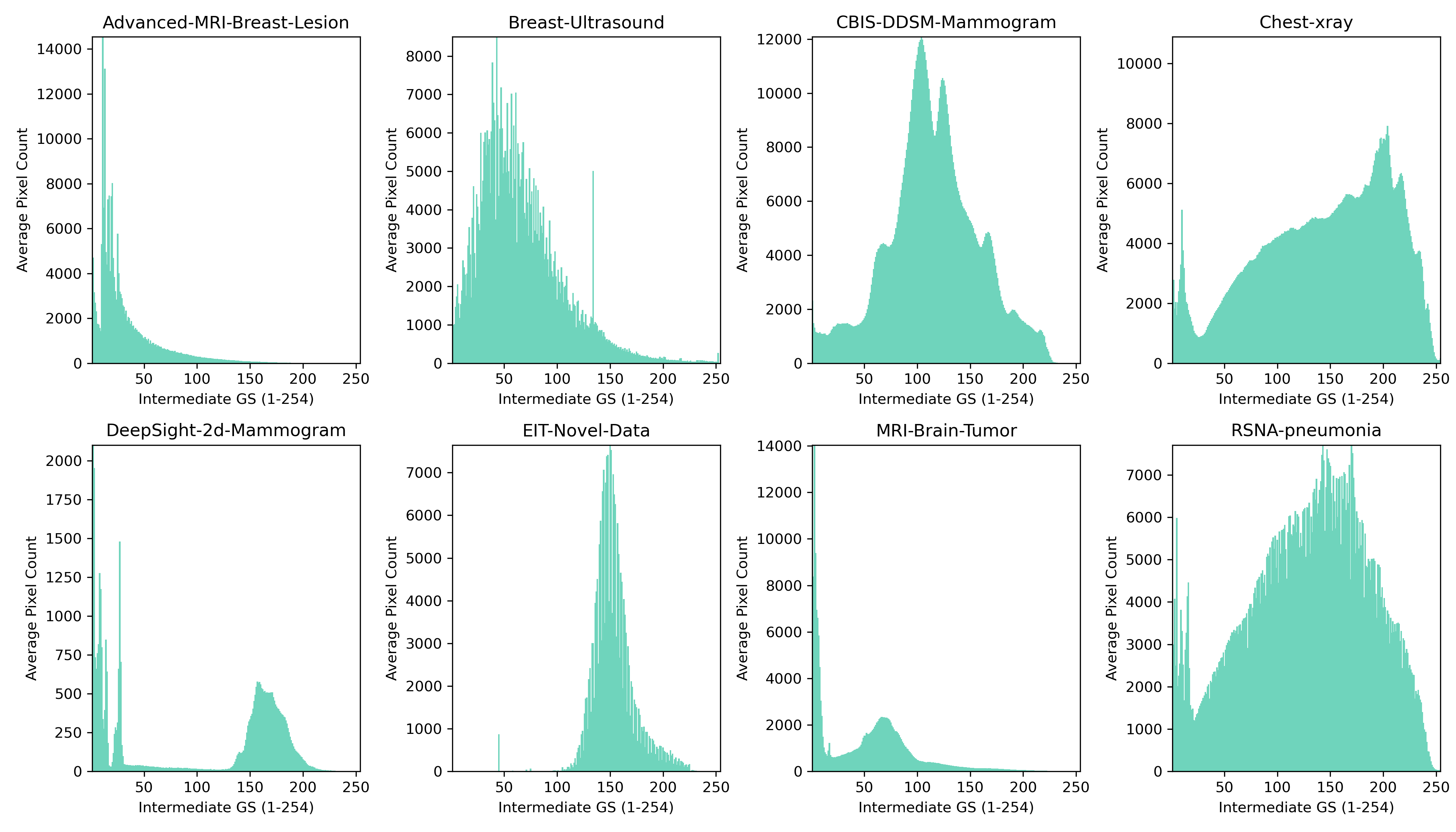
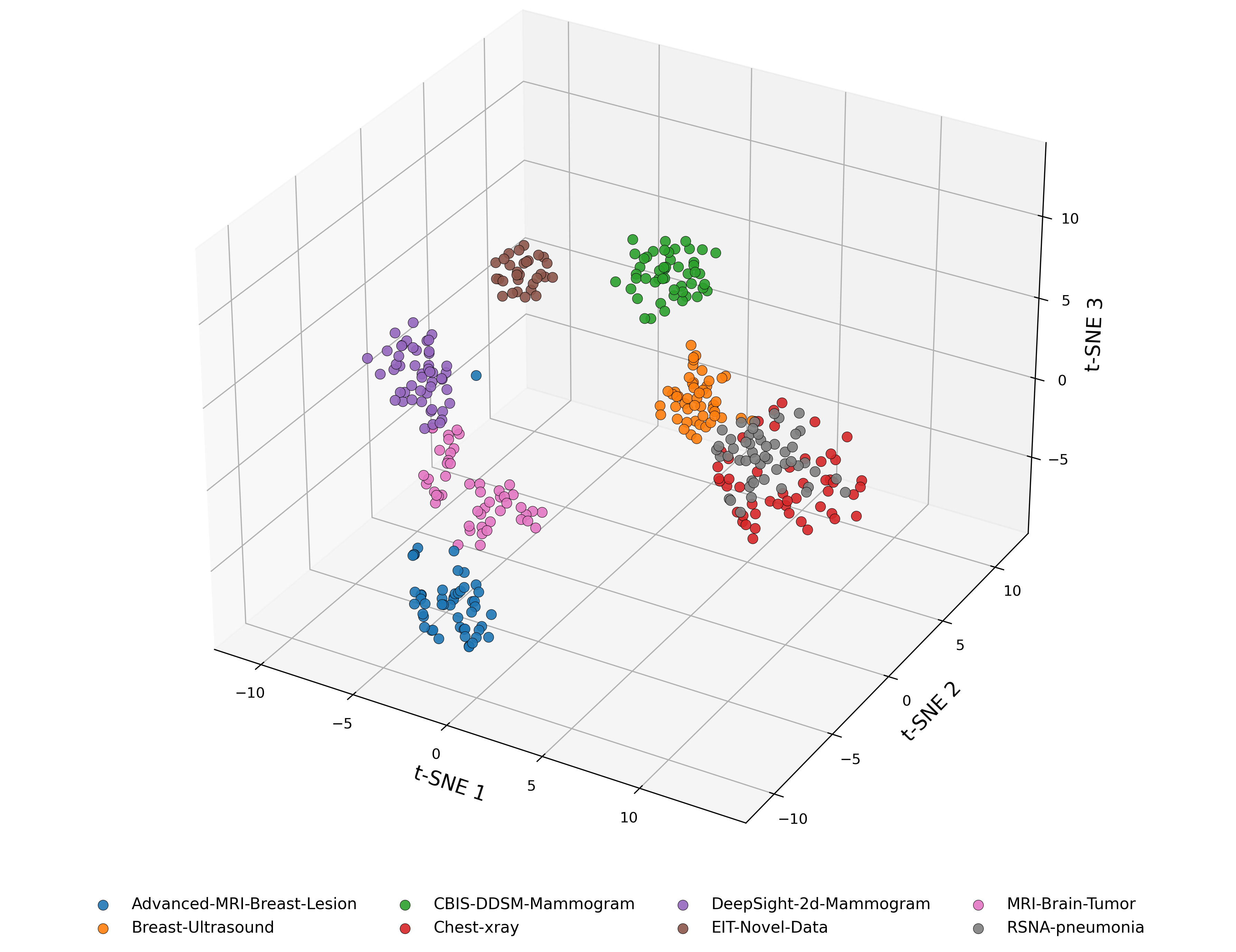
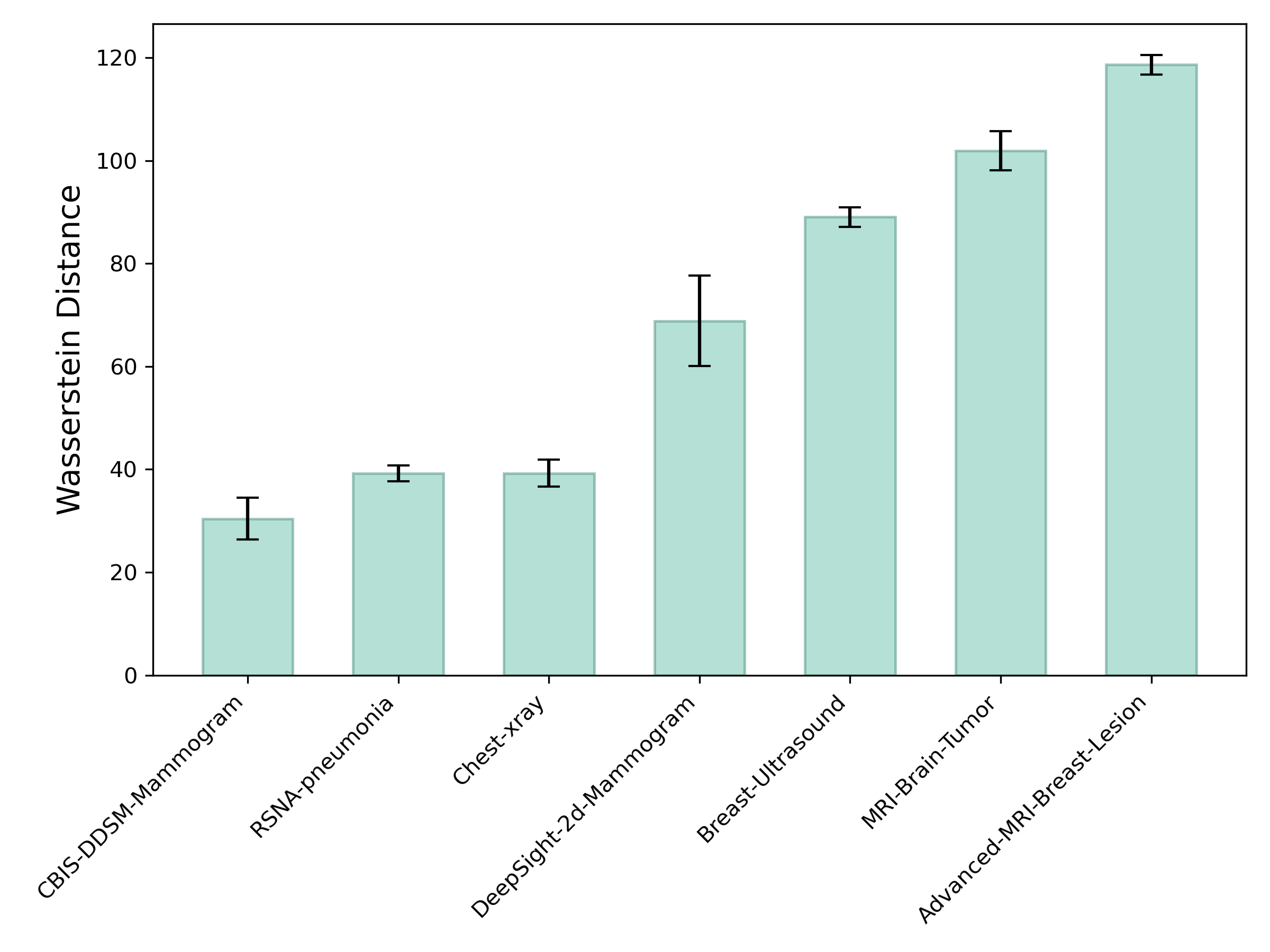
The t-SNE visualization in Figure 3a reveals that each dataset forms a distinct cluster in the CNN embedding space, highlighting clear modality-specific differences. The EIT-Novel-Data cluster is positioned relatively close to most of the datasets, suggesting that it shares feature-level similarity across multiple modalities. However, it's more clearly separated from the Advanced-MRI-Breast-Lesion and chest X-ray datasets. This pattern partially aligns with the Wasserstein analysis since EIT is close to mammography in both views, but despite intensity-based similarity to X-ray, the embedding places EIT farther from the X-ray datasets, indicating that relying on grayscale similarity alone is insufficient when determining similarity in representational structures.
We hypothesize that training a meta-model using datasets that are proximally-close to the target EIT dataset in the embedding space will yield the best model performance. This is because meta-learning involves learning to learn from limited data, and proximally-related tasks share similar representational structures, thus providing a better model initialization for rapid adaptation to the target domain.
Data Processing Pipeline
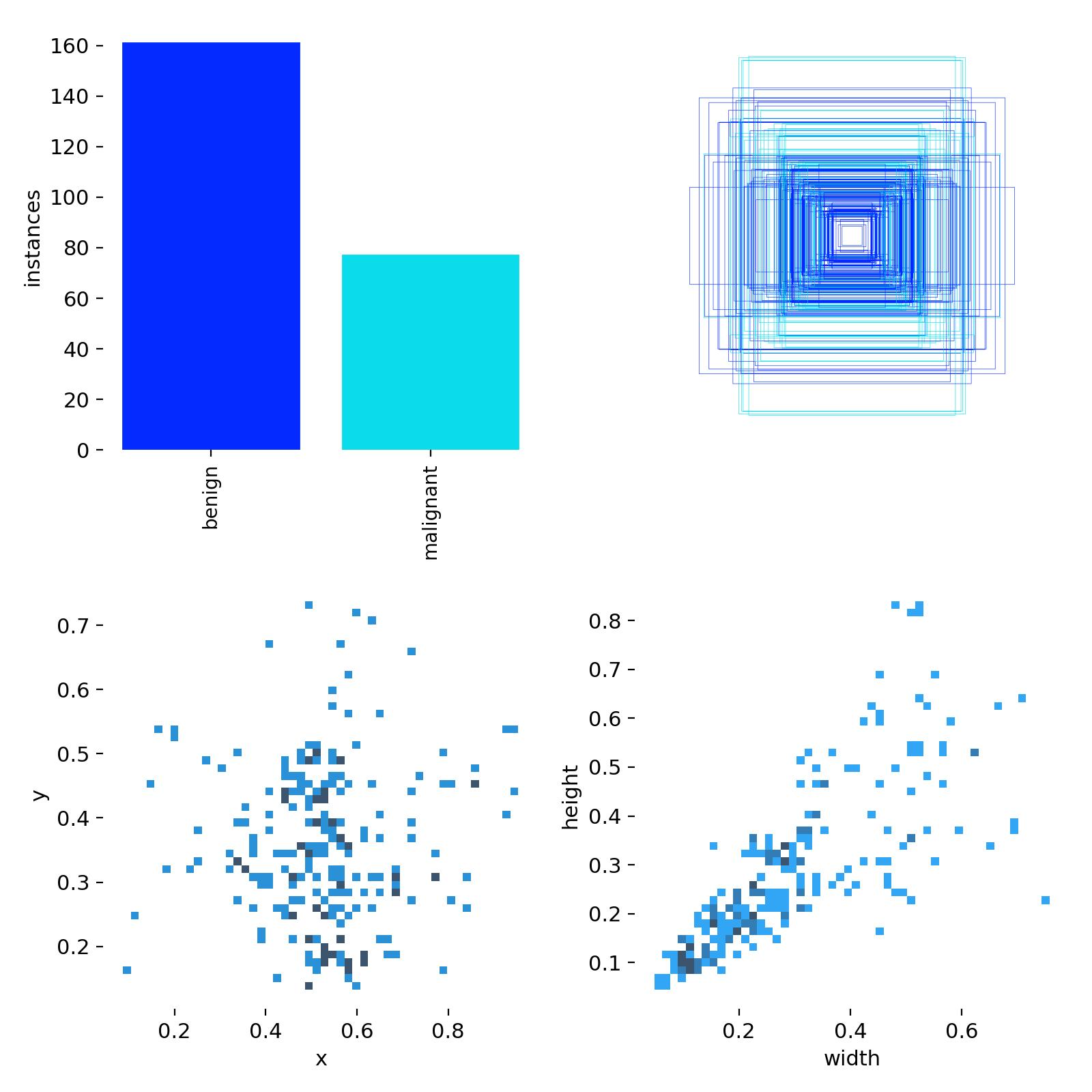
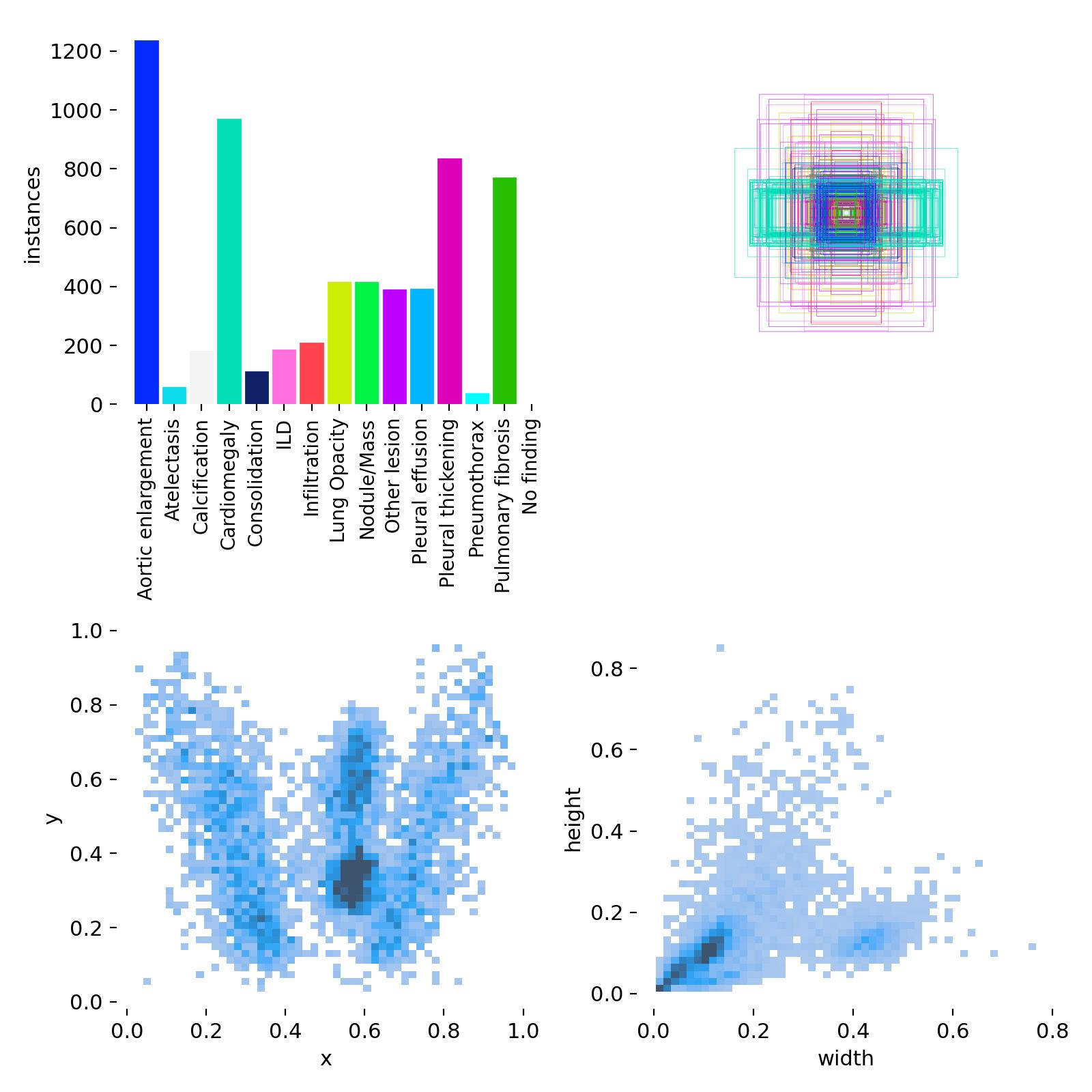
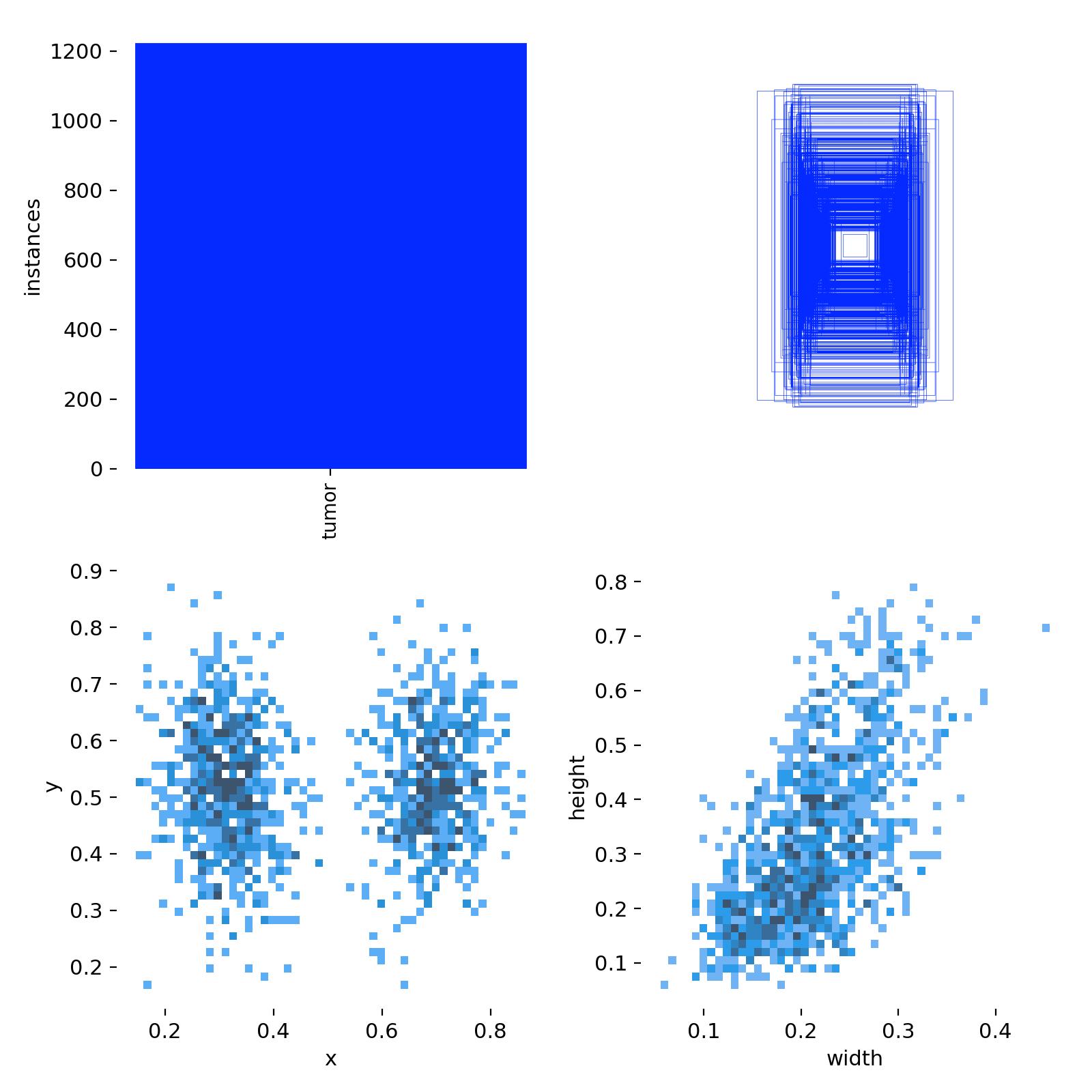
Following ingestion, datasets underwent a standardized cleaning and transformation workflow. All images were inspected for corruption by verifying that the headers and metadata were readable during decompression, and ensuring that each image could be successfully parsed by standard imaging libraries e.g., pydicom, PIL, OpenCV. Files that failed these checks were excluded, and datasets were further screened for outliers such as images with anomalous dimensions, empty pixel arrays, or intensity distributions inconsistent with the rest of the dataset. Any images that did not have corresponding annotation files, or vice versa, were discarded. Metadata associated with each dataset was normalized into a unified schema, allowing consistent representation of imaging modality, pathology, and annotation type, while removing unique identifiers to maintain patient anonymity. Preprocessing steps included contrast normalization, 8-bit grayscale conversion, and resizing of images to meet the requirements of downstream models. The system is capable of handling imaging data in multiple formats, including DICOM, JPG, PNG, HDF5, JPEG, MAT and NPZ. Since the datasets spanned both detection and segmentation tasks, i.e., images were accompanied with either bounding boxes or segmentation masks, annotation harmonization was performed to establish a consistent format to enable objection detection across multiple modalities. In particular, segmentation masks were converted into bounding box representations, and coordinate systems and bounding box annotations were standardized to ensure interoperability across datasets. Sanity checks using an image-label overlay routine were performed post-harmonization to ensure the transformed bounding box labels preserved their precise location relative to the anomalies in the image. For evaluation purposes, datasets lacking predefined splits were partitioned into training and validation sets using an 80/20 ratio.
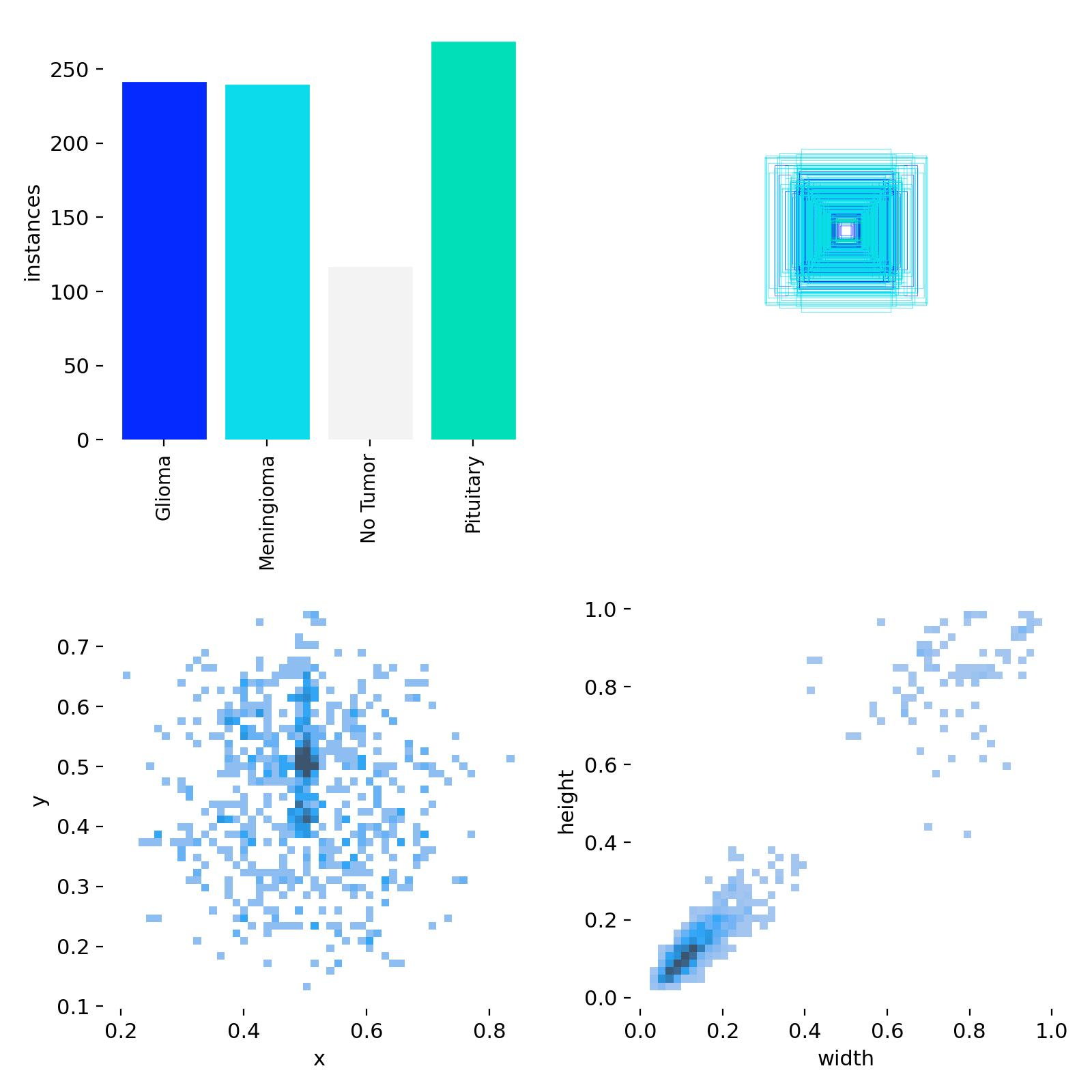
To support flexible workflows, the system includes an editable pipeline-config file that allows users to specify which datasets to ingest, process, and store, as well as which datasets to feed to the training pipeline, enabling either selective handling of individual datasets or processing of the entire portfolio. To optimize memory and compute, the pipeline automatically skips datasets that have already been processed, while providing users the option to force re-processing when needed. Through the above methods, the pipeline enables the scalable reuse of large, heterogeneous datasets in a manner that ensures consistency and reproducibility. The resulting standardized datasets serve as high-quality inputs for meta-learning models, providing a foundation for evaluating few-shot detection on novel imaging tasks.
Meta Learning Algorithm
Meta Training
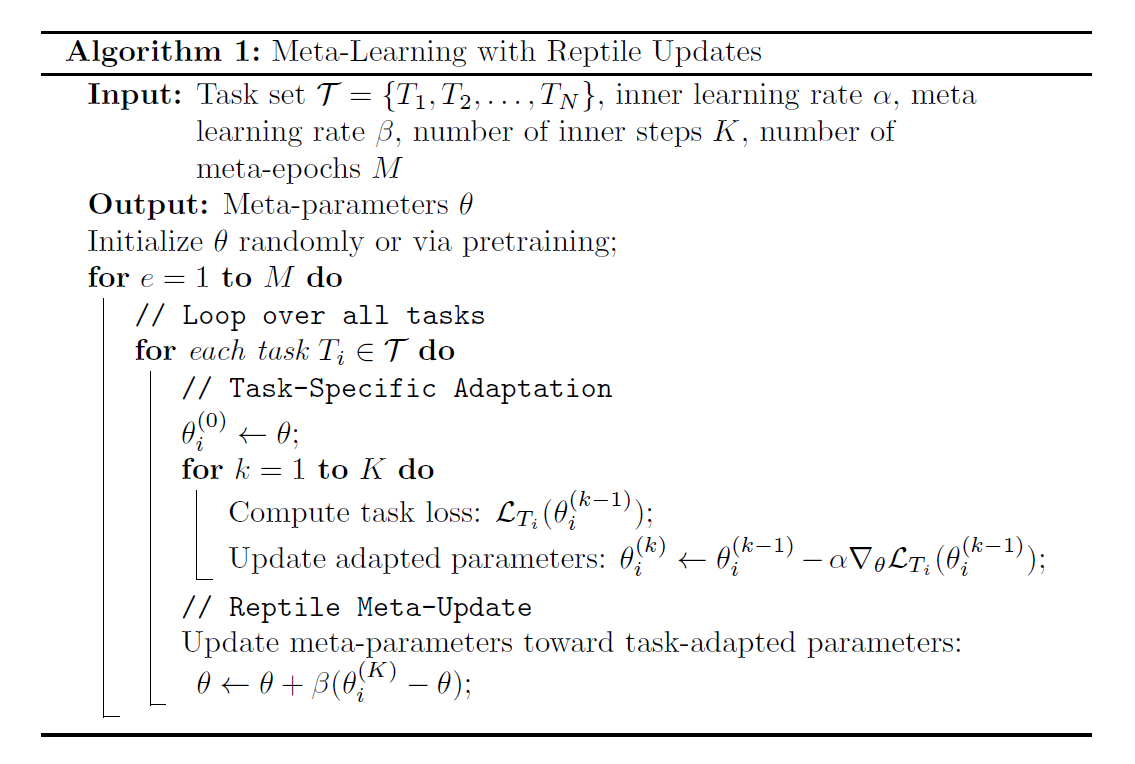

Meta Validation
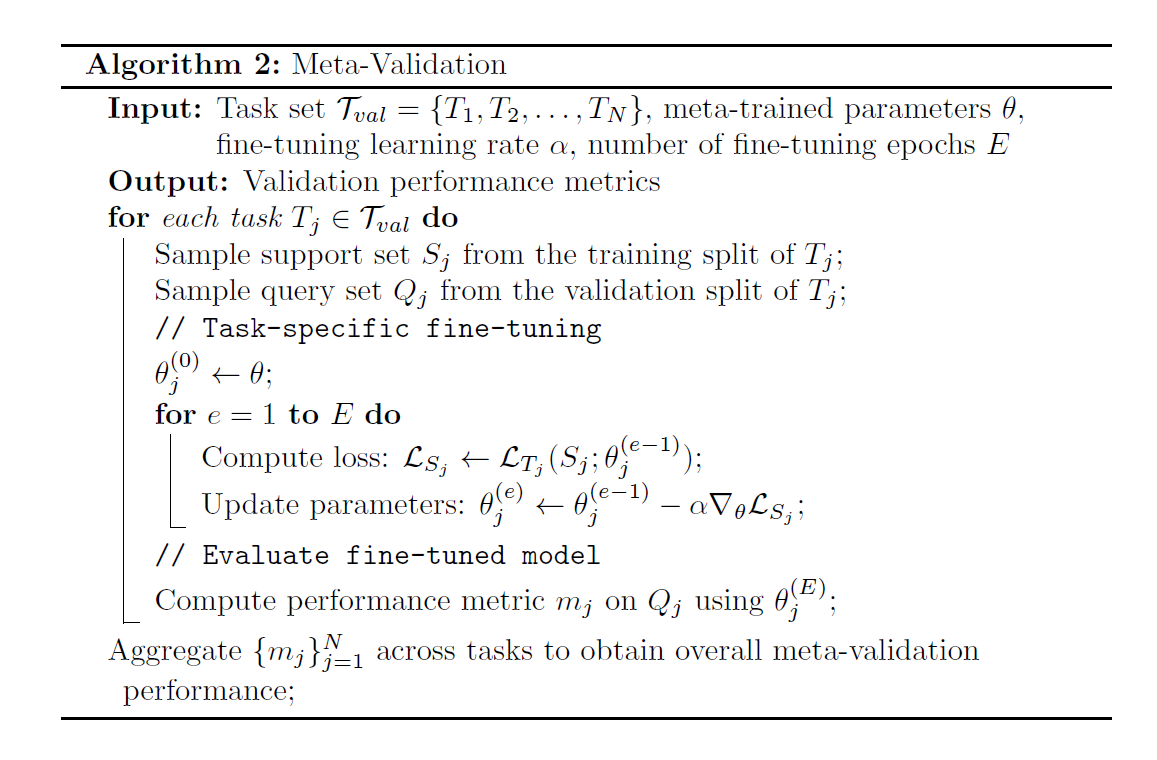

Few-Shot Learning
One can go a step further and incorporate the new task into the meta-model by including it as an additional task during meta-training. This can be achieved with ease because the meta-learning algorithm is designed to extend to new tasks without needing to retrain the model from scratch. Incorporating the new task nudges the meta-parameters toward an optimal minima that can quickly adapt to tasks that share similarities with the new task. This strengthens the model's long term adaptability as more tasks are introduced in the future, improves robustness by reducing the need for extensive fine-tuning, shortens deployment timelines, and ensures the model can continuously accommodate process or domain shifts without sacrificing performance on existing tasks.
Meta-Model Setup
The following model parameters were used for training and evaluation of the meta-learning model.
Table 3: Meta-Model Parameters
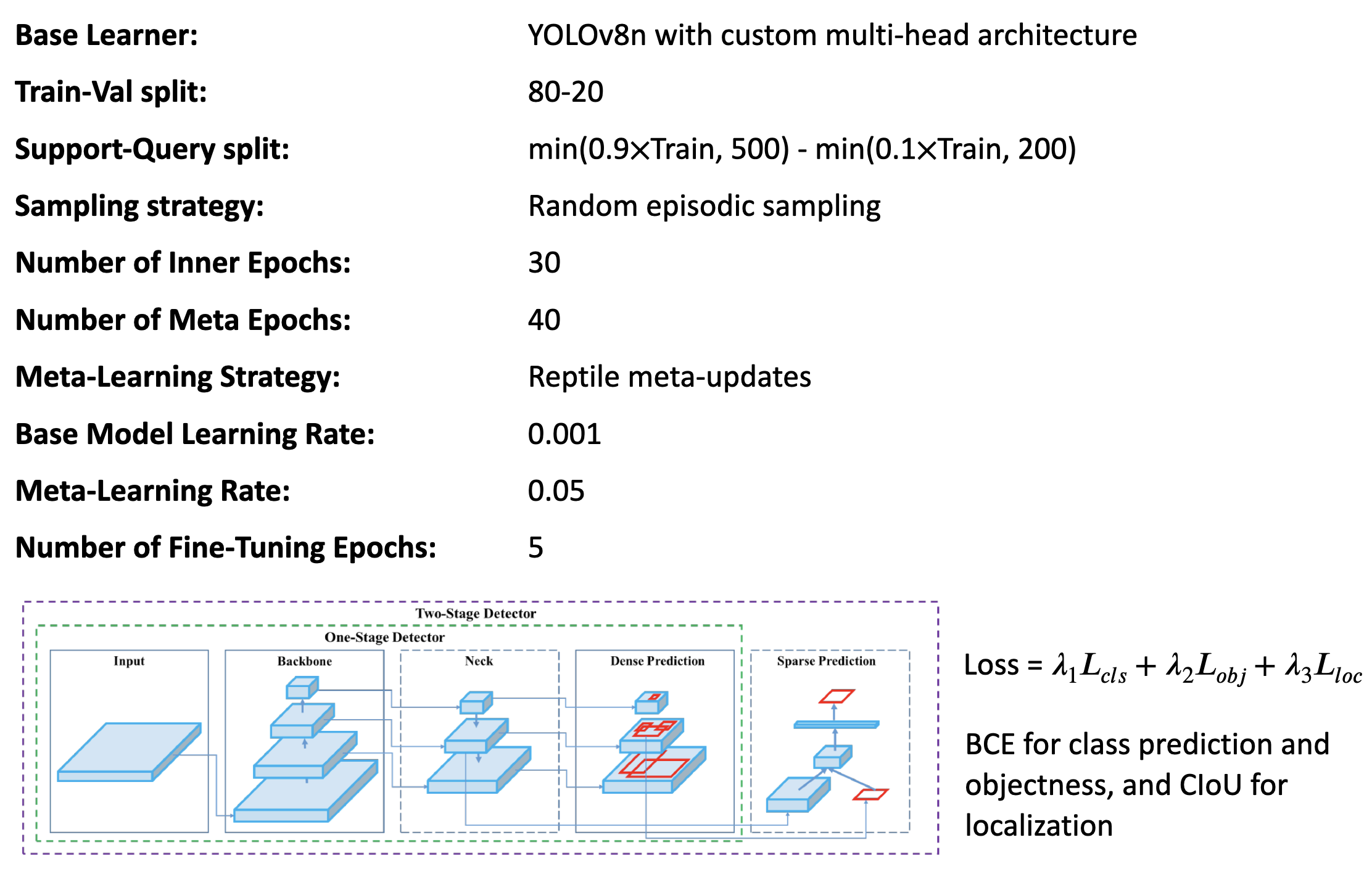
Experiments & Findings
We conducted ablation studies to identify optimal model hyperparameters, quantify how the number and combination of tasks spanning different modalities affect meta-learning generalizability, assess the impact of base learner architecture on accuracy, evaluate meta-model performance under extreme few-shot conditions, and determine the minimum fine-tuning required to maintain high accuracy.
Experiment 1 - Optimal Meta-Learning Rate
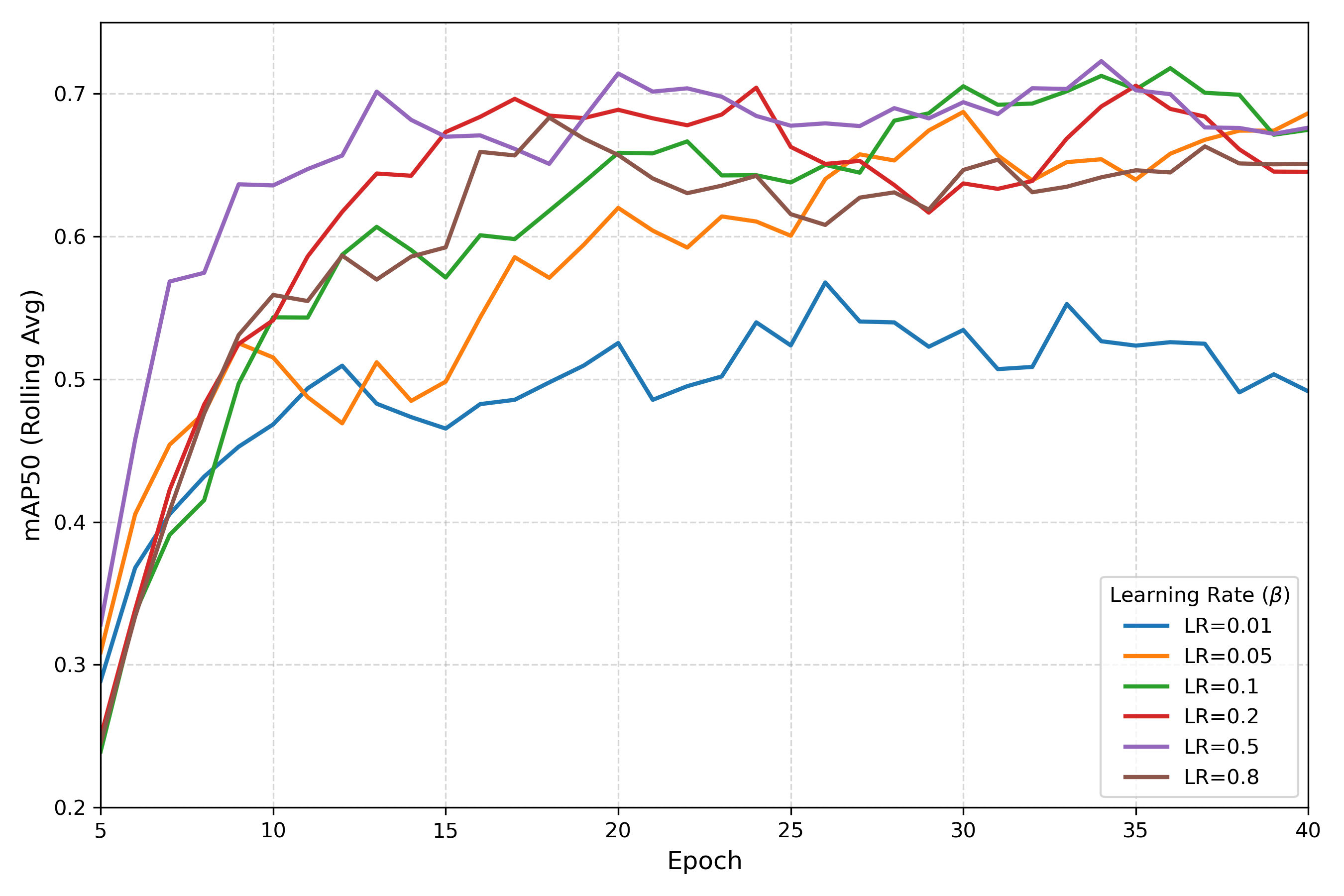
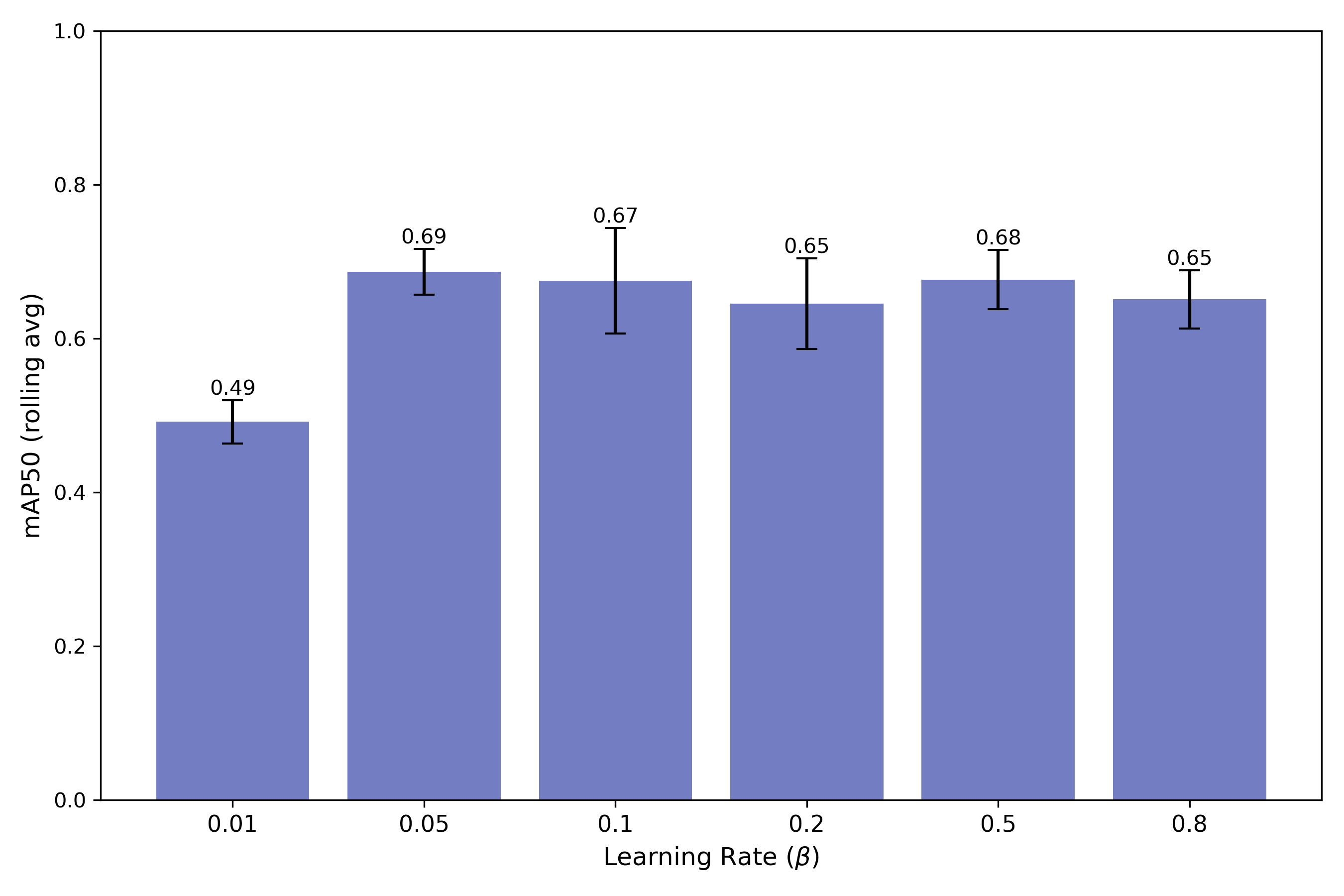
Experiment 2 - Generalizability and Task Dependence
The model's performance trajectory supports the hypothesis that meta-model generalization benefits from increased task diversity up to an optimal point. The initial improvement going from 2 to 5 tasks suggests that exposure to more diverse medical imaging tasks enhances the meta-learner's ability to quickly adapt to new detection problems by learning more generalizable feature representations. Beyond 5 tasks, the meta-model's generalizability begins to deteriorate with the inclusion of out-of-distribution chest X-ray tasks. This is due to task interference introducing negative transfer effects that counter the benefits of diversity. The performance degradation is most pronounced in precision, suggesting that discriminative capabilities become compromised when the meta-learner attempts to accommodate too many disparate visual domains and detection requirements simultaneously.
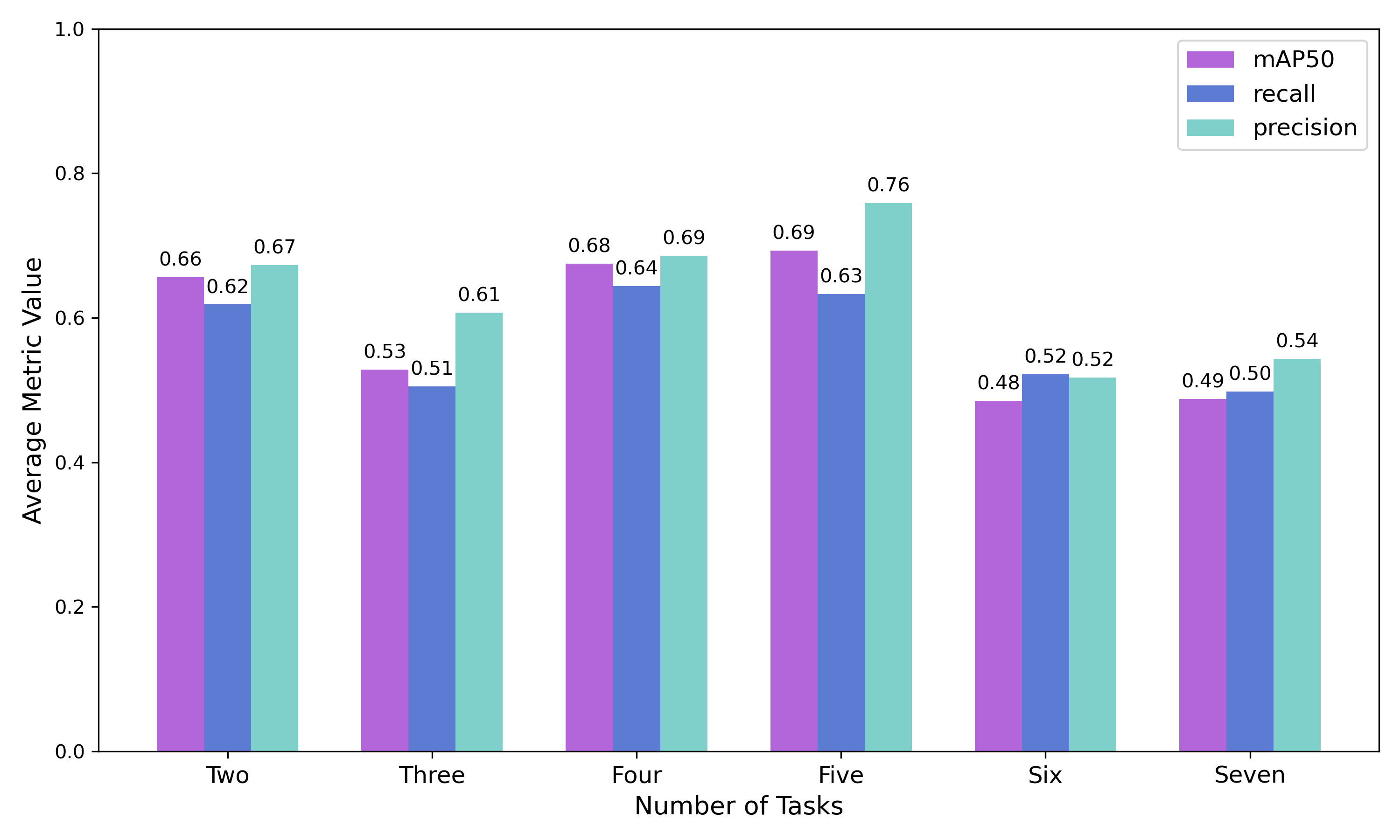
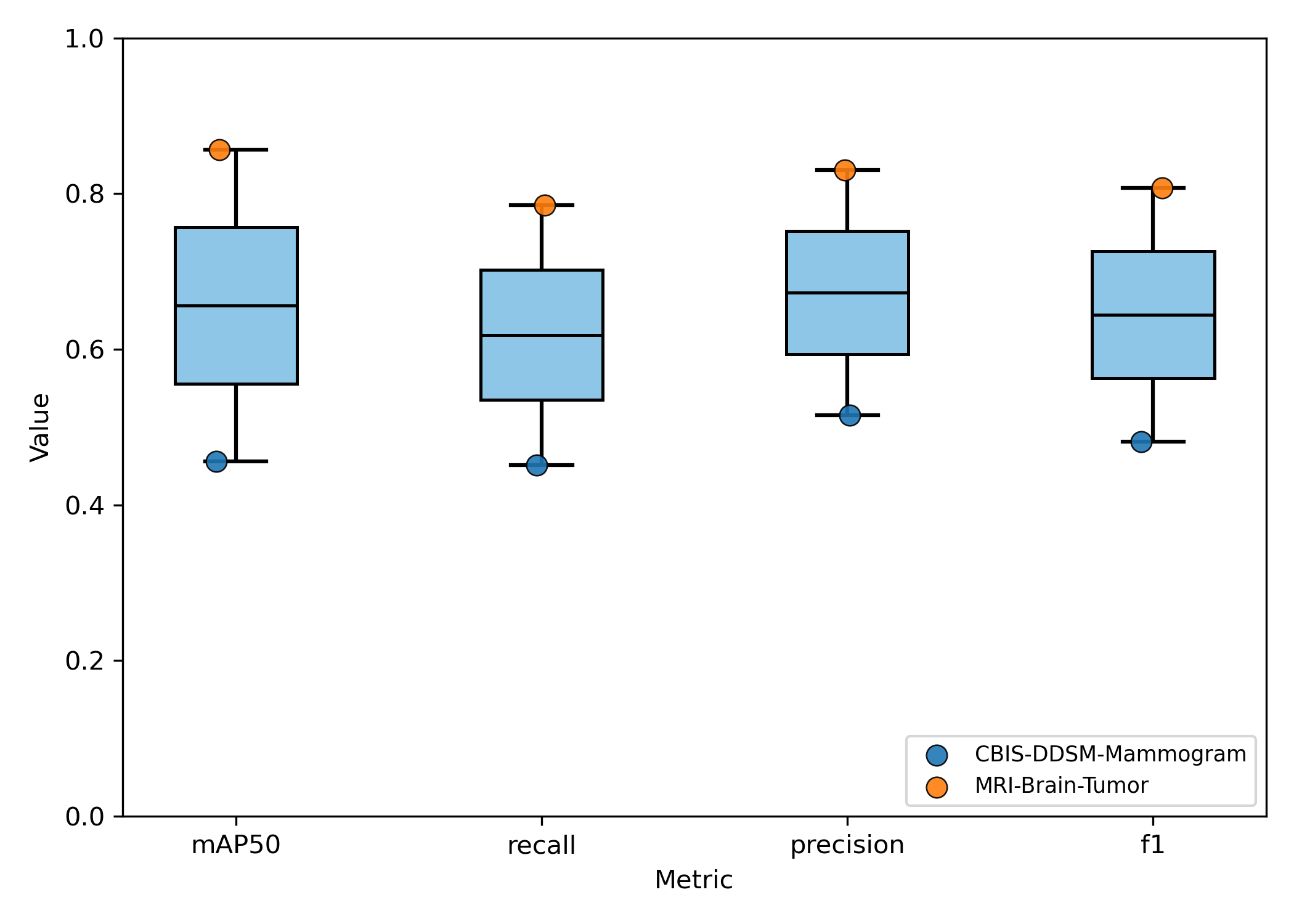
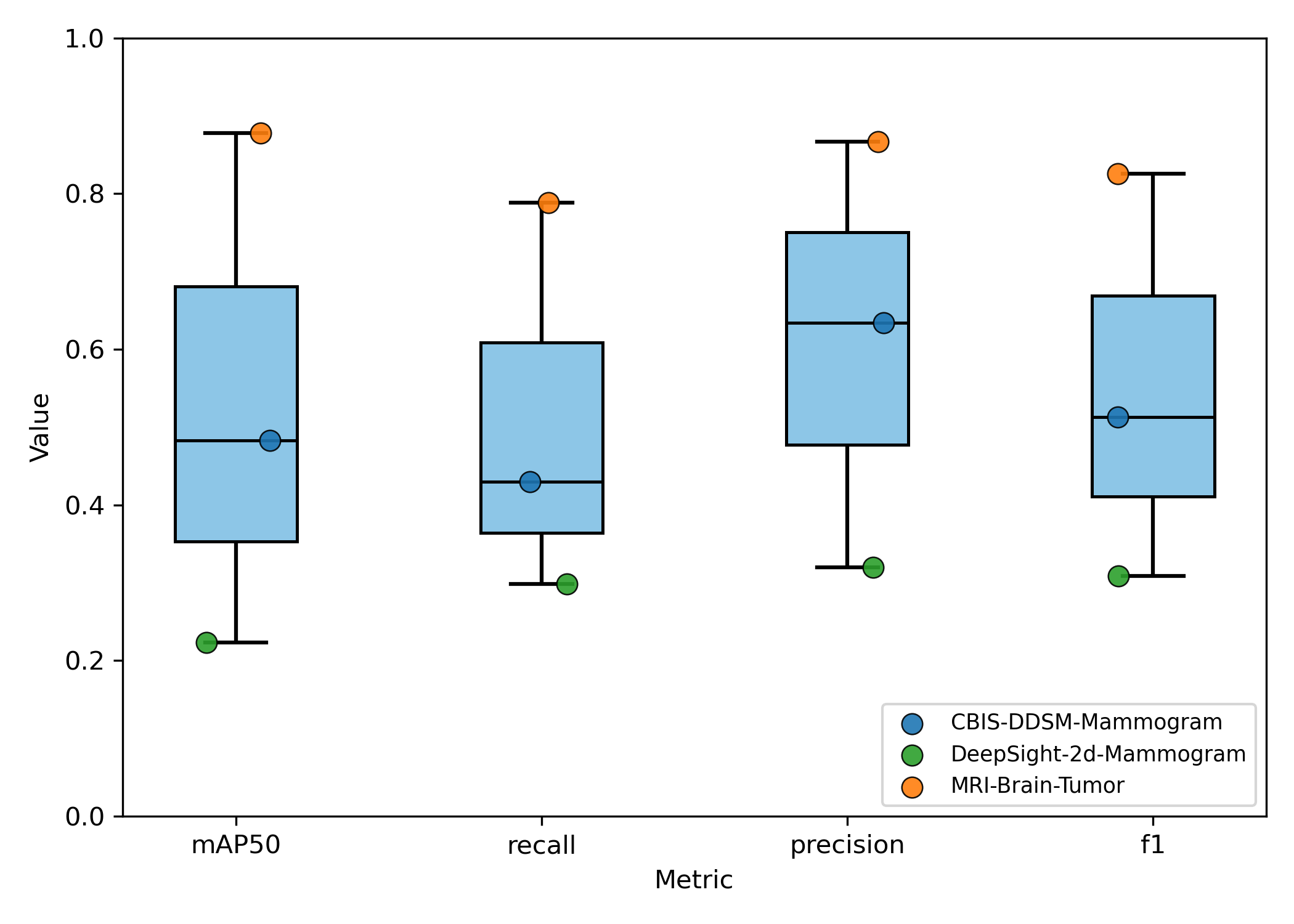
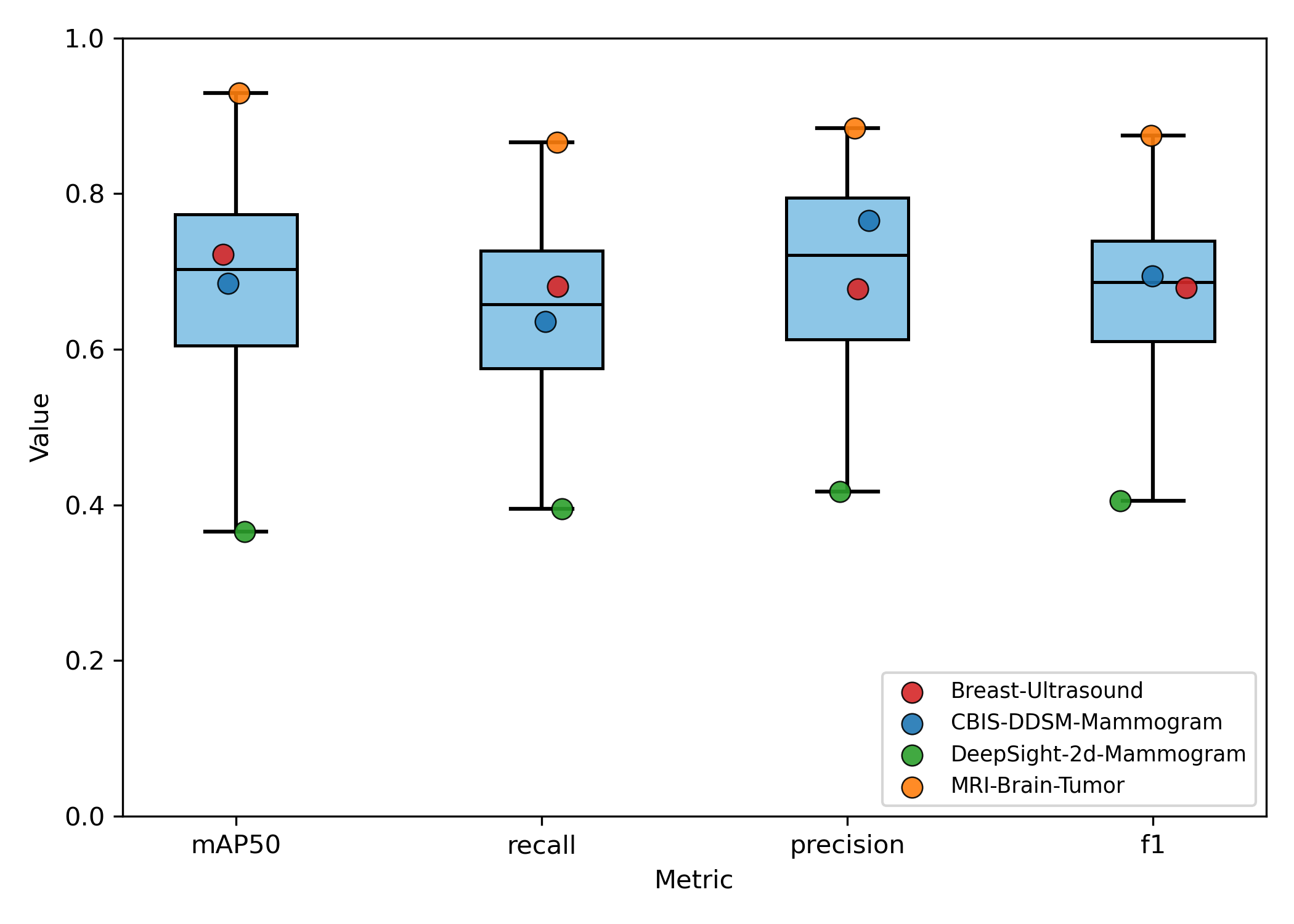
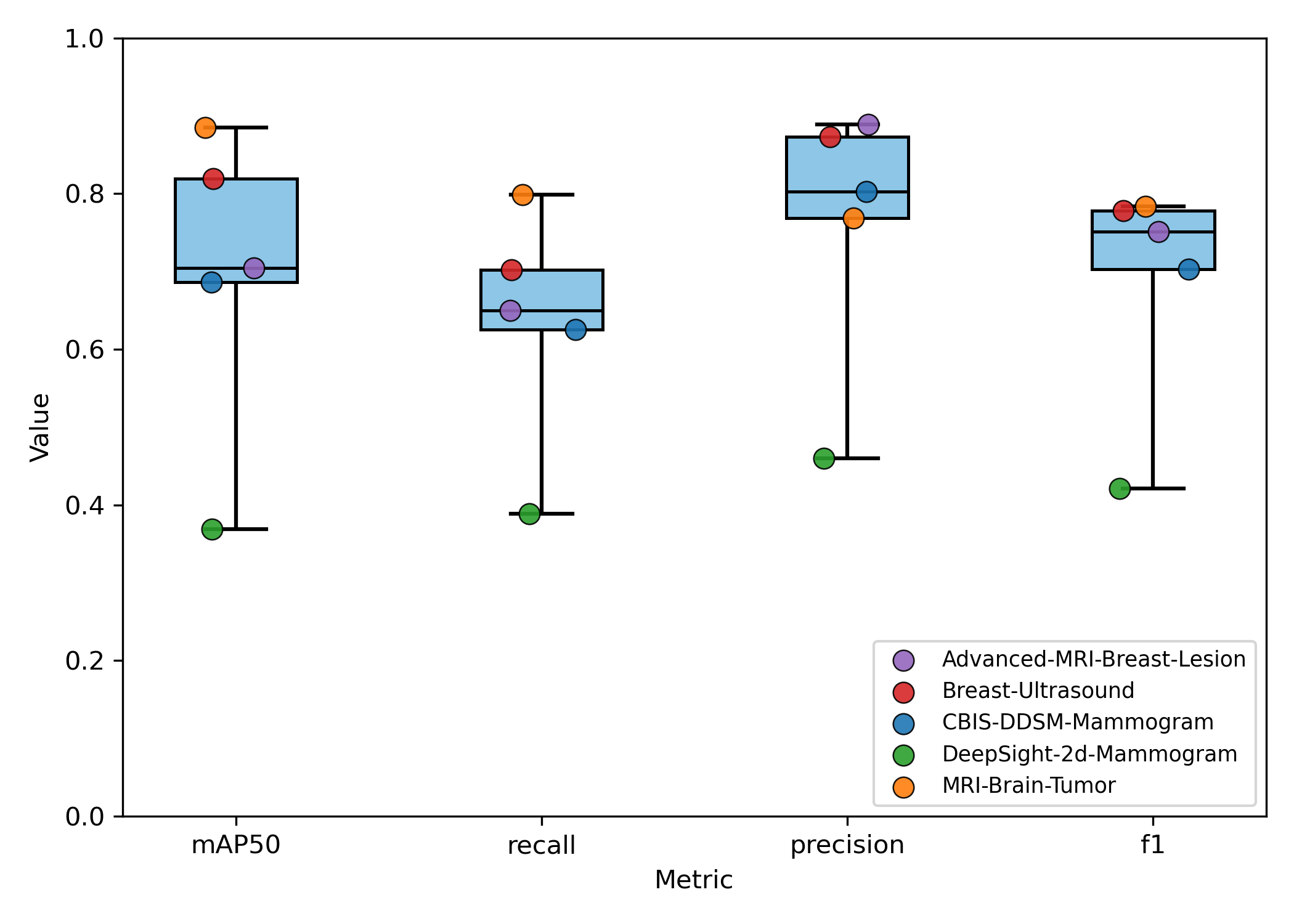
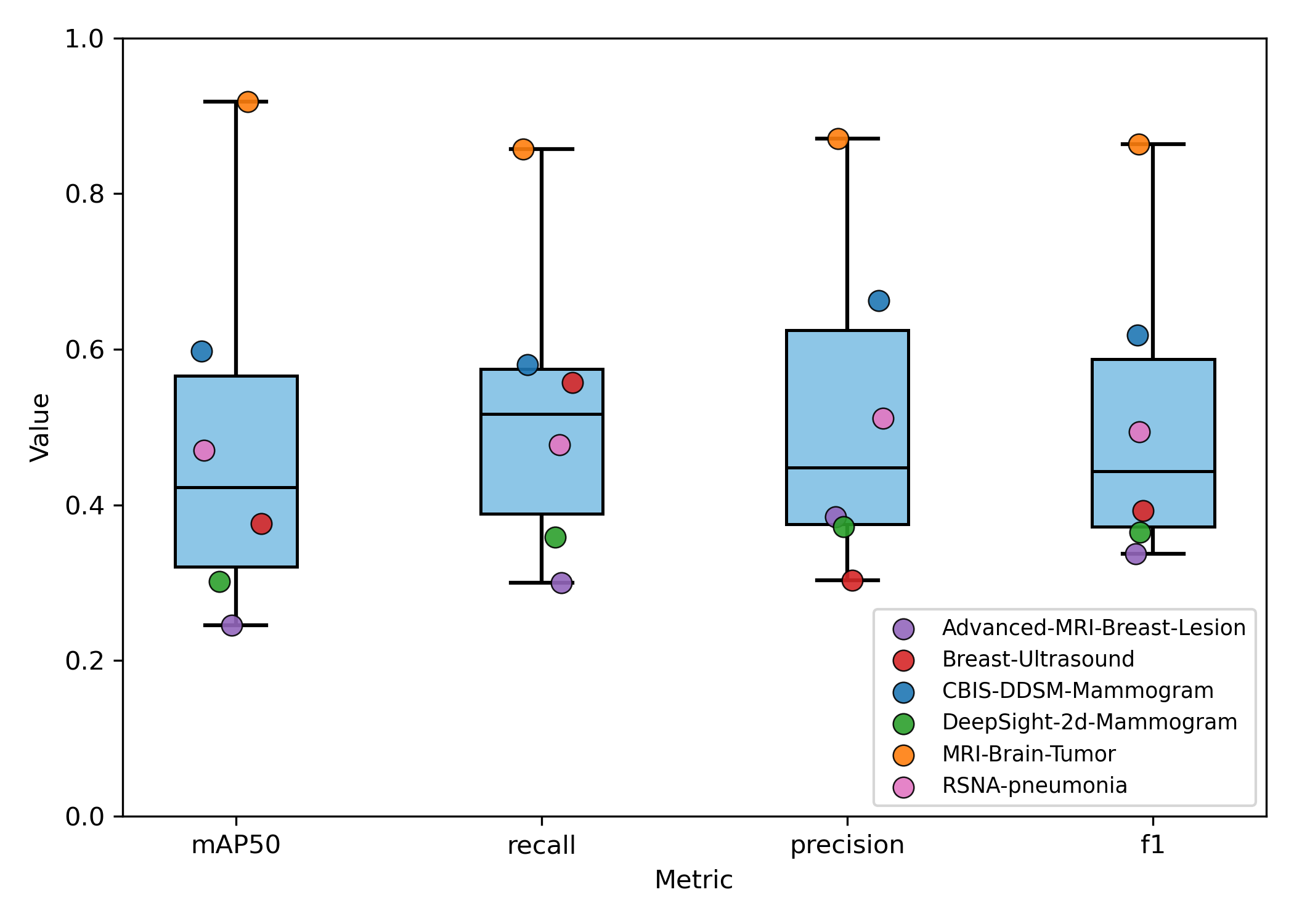
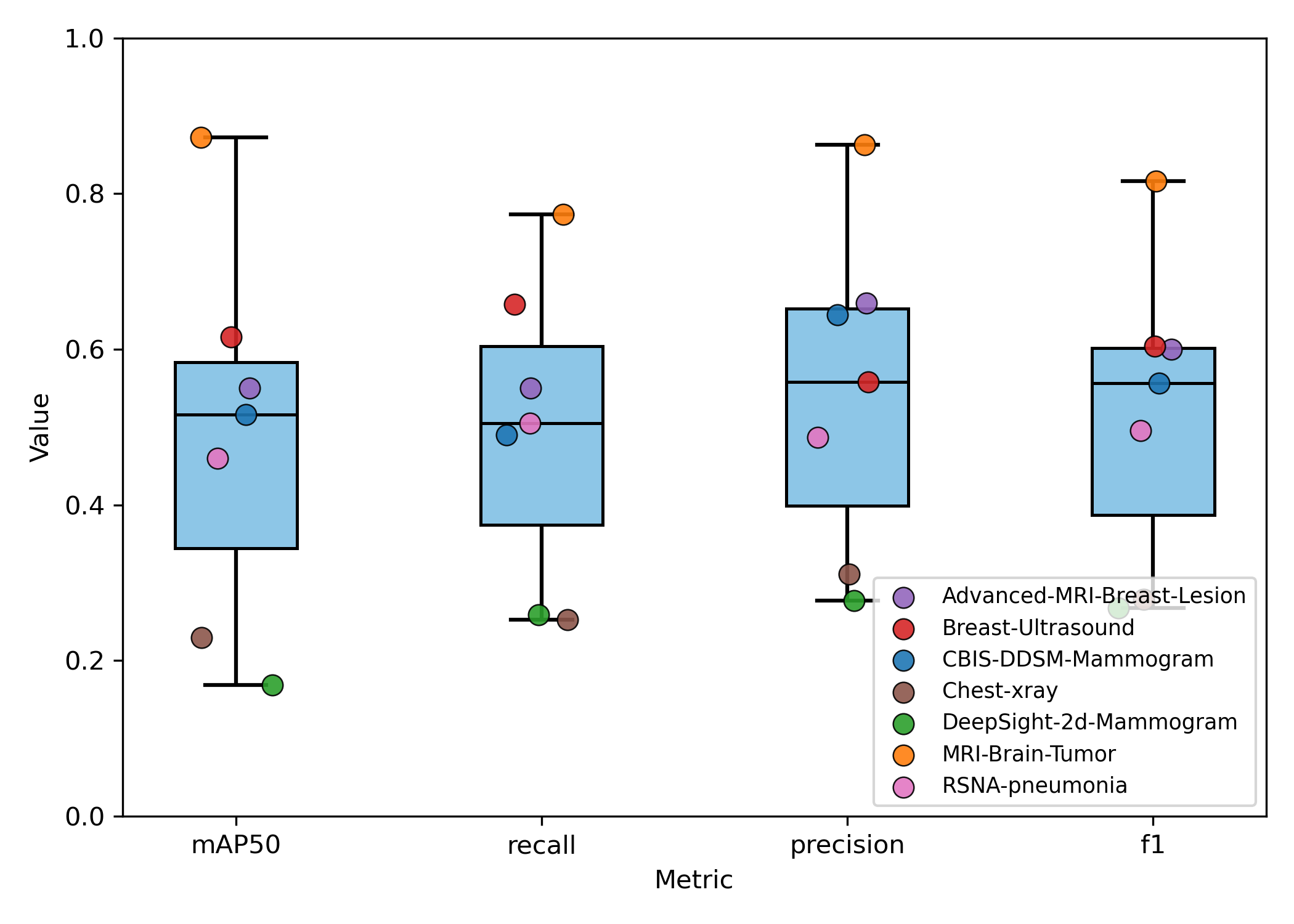
These findings indicate an optimal selection of tasks with sufficient heterogeneity to improve generalization without causing destructive interference to the meta-model's learned parameters.
Experiment 3 - Multi-task Meta-Learning Performance
In experiment 3, we investigated the impact of fine-tuning schedules on meta-model performance. As expected, the model showed improved performance by 4-27% across tasks when the number of fine-tuning steps was increased from 5 to 20. By doing more gradient descent steps before making a prediction on a given task, we are moving the general meta-model’s parameters closer to the optima of the task at hand, which results in better performance.
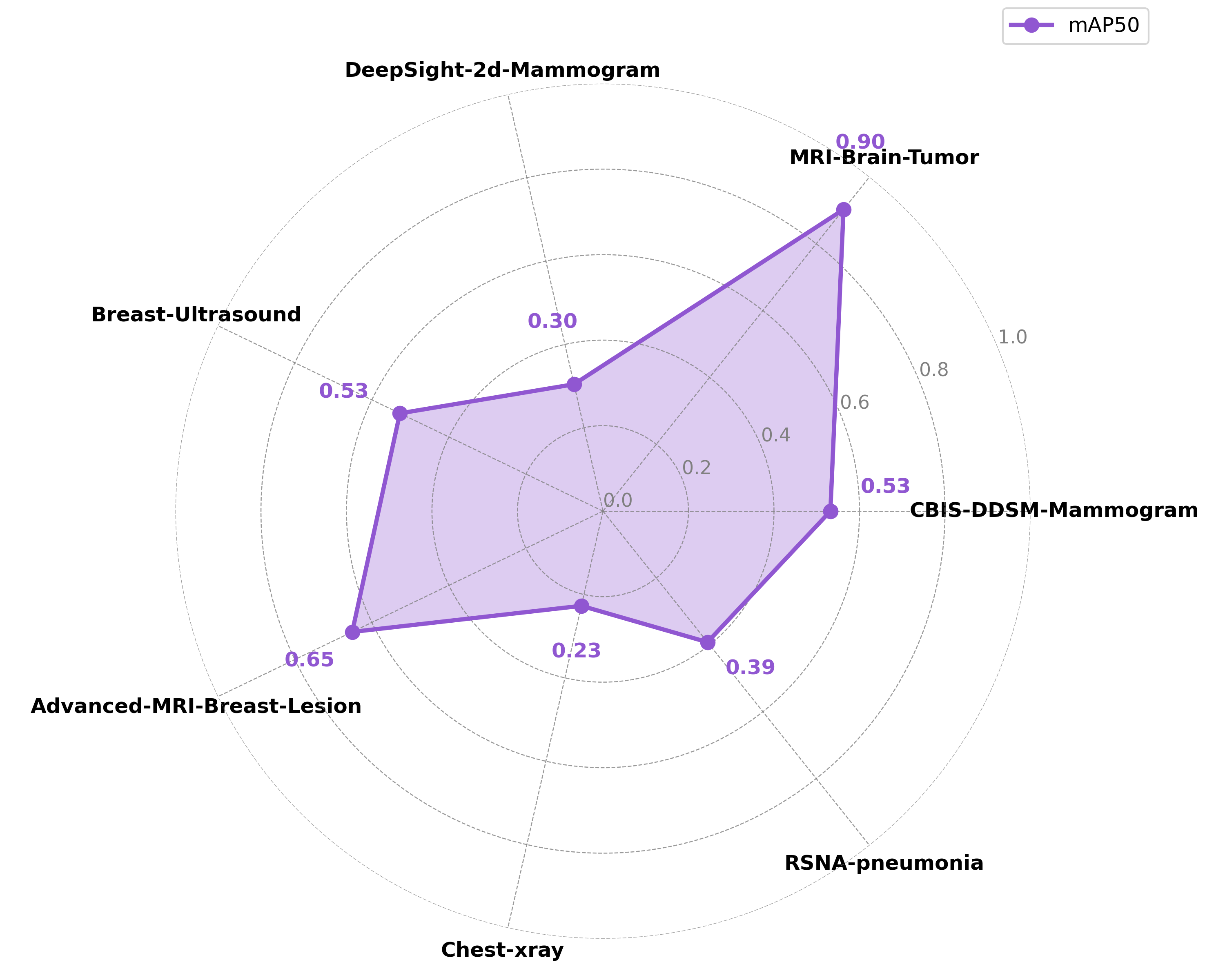
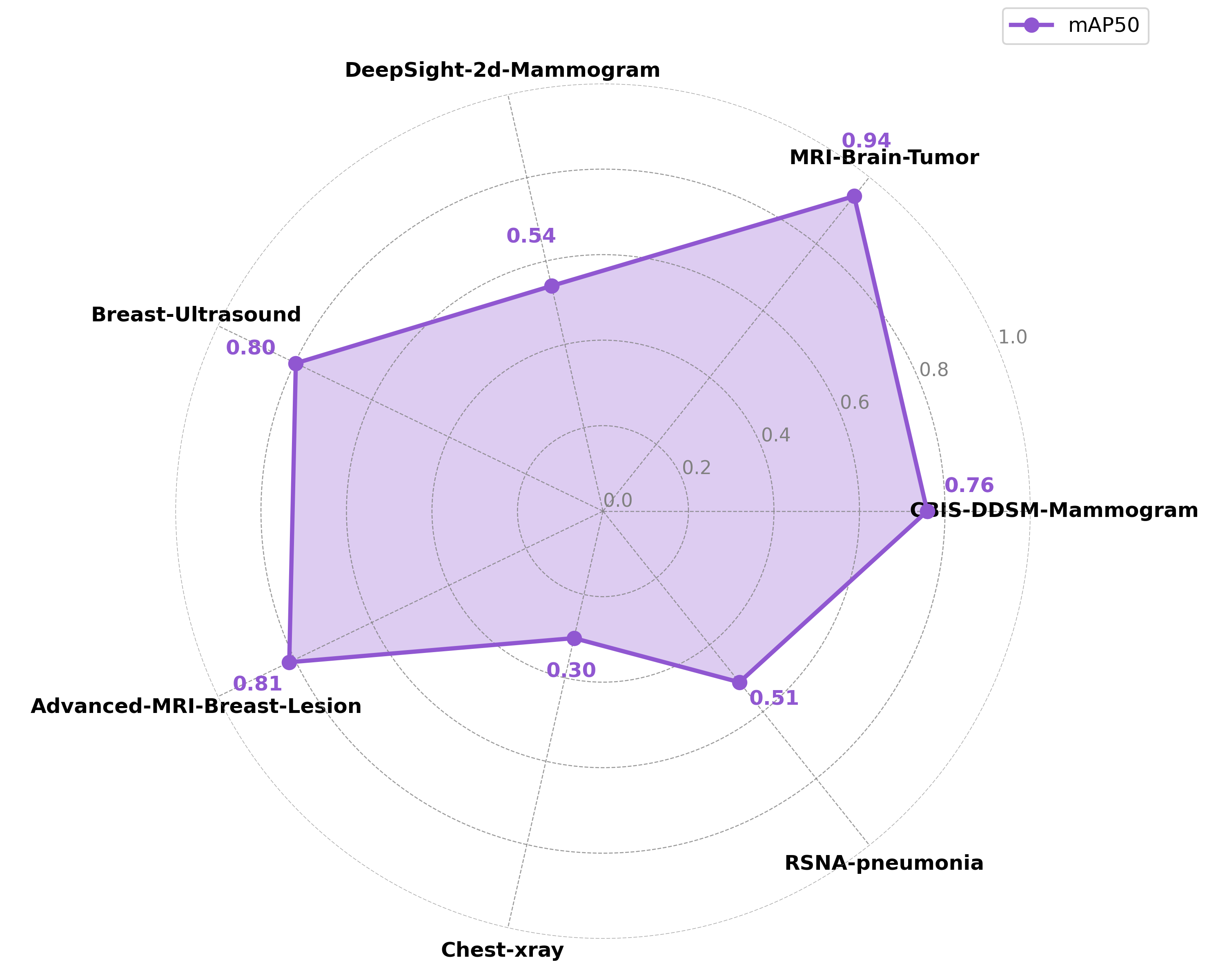
Experiment 4 - Model Size and Warmup Schedule
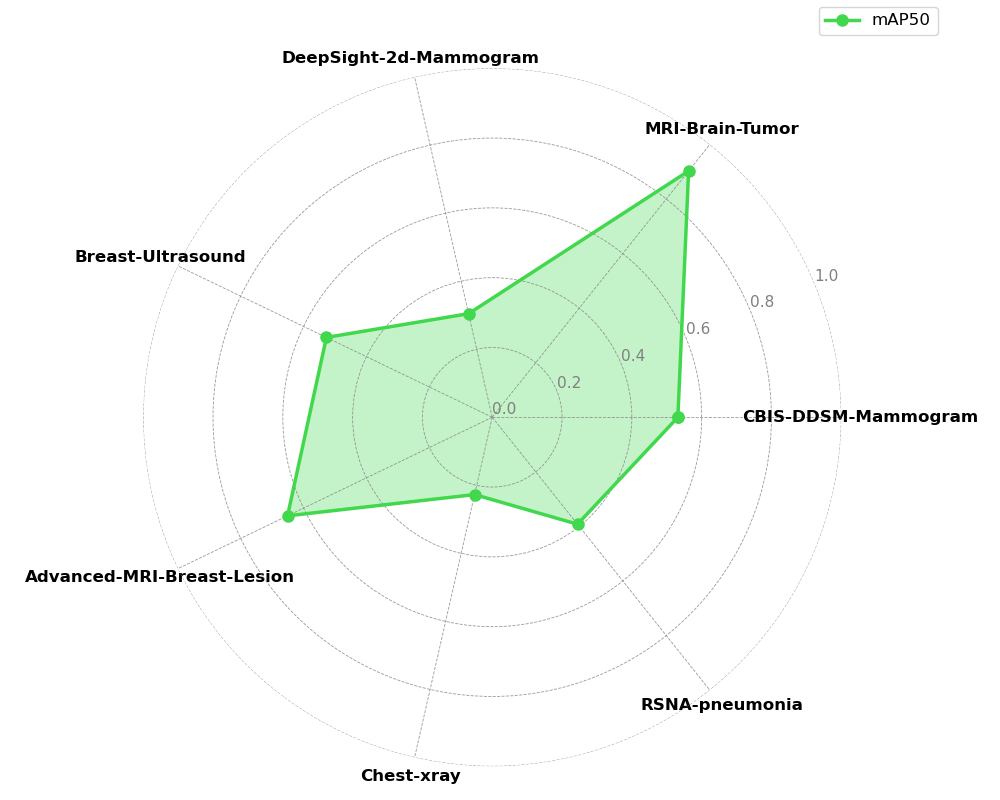
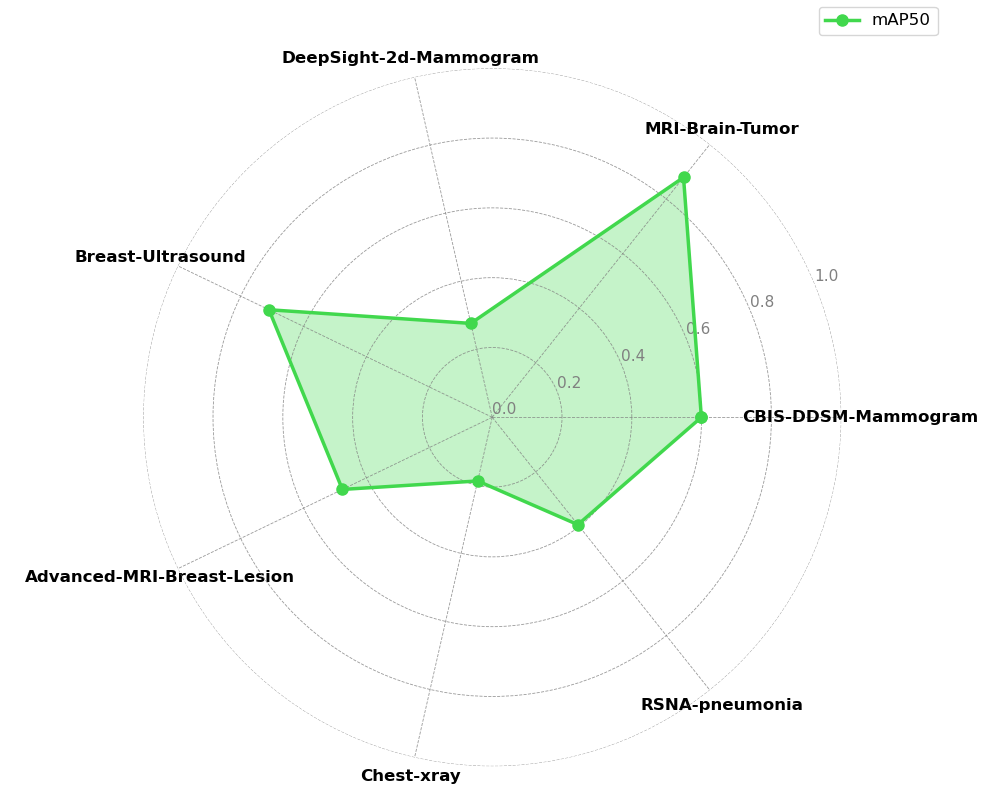
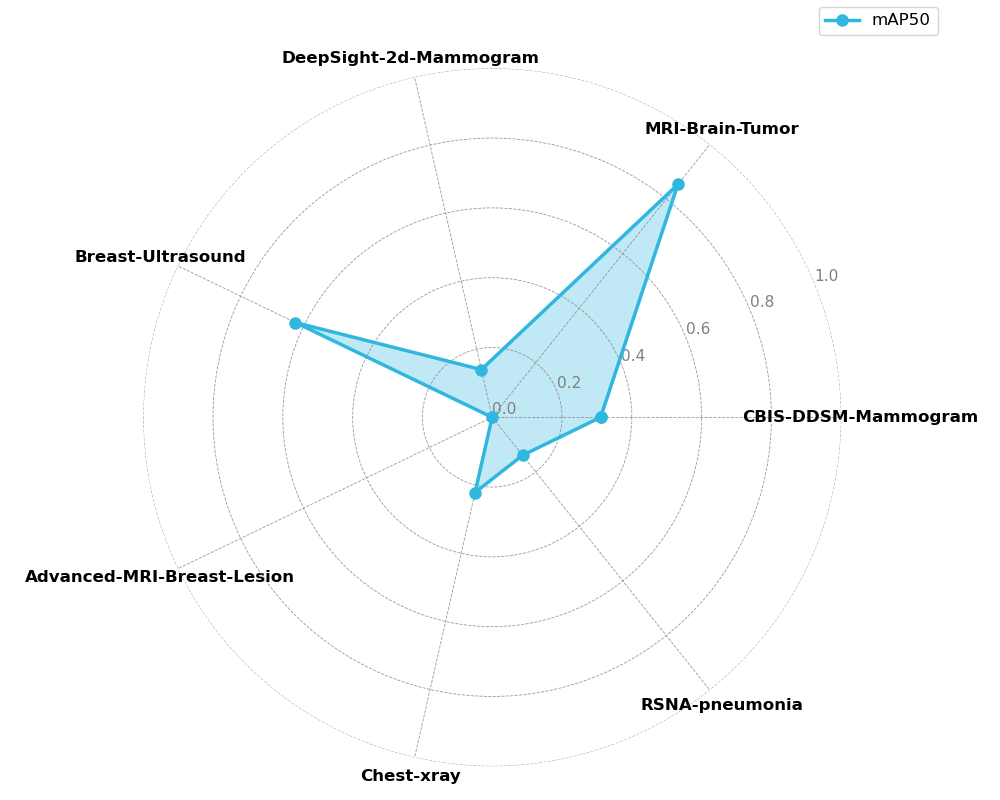
Experiment 5 - Comparison with Incremental Transfer Learning
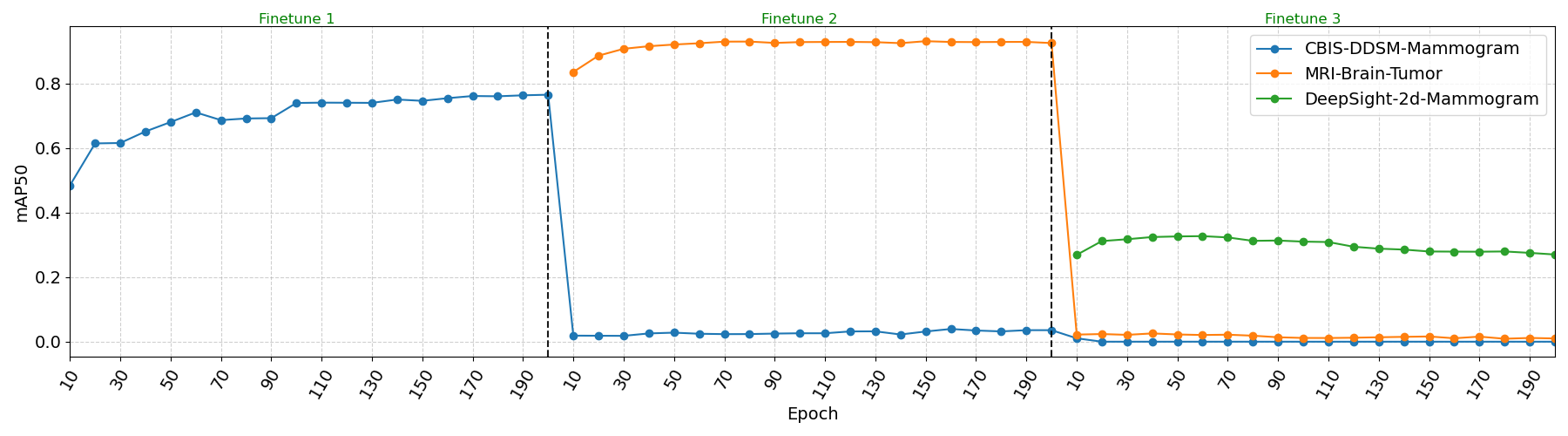
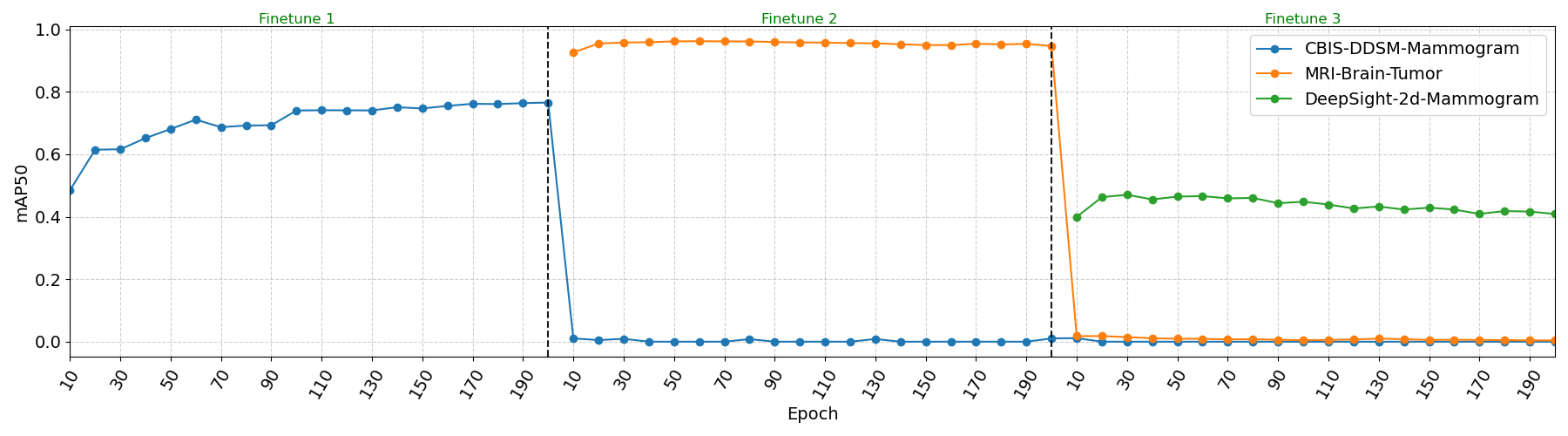
The incremental transfer learning results reveal severe limitations inherent to this approach. The line plots demonstrate clear evidence of catastrophic forgetting, where previously learned tasks experience immediate and severe performance degradation to near-zero mAP50 when the model begins learning subsequent tasks. These results show that traditional fine-tuning approaches cannot maintain multi-task competency in medical imaging domains, and are especially unfit for use cases that require new datasets to be integrated in the training pipeline.
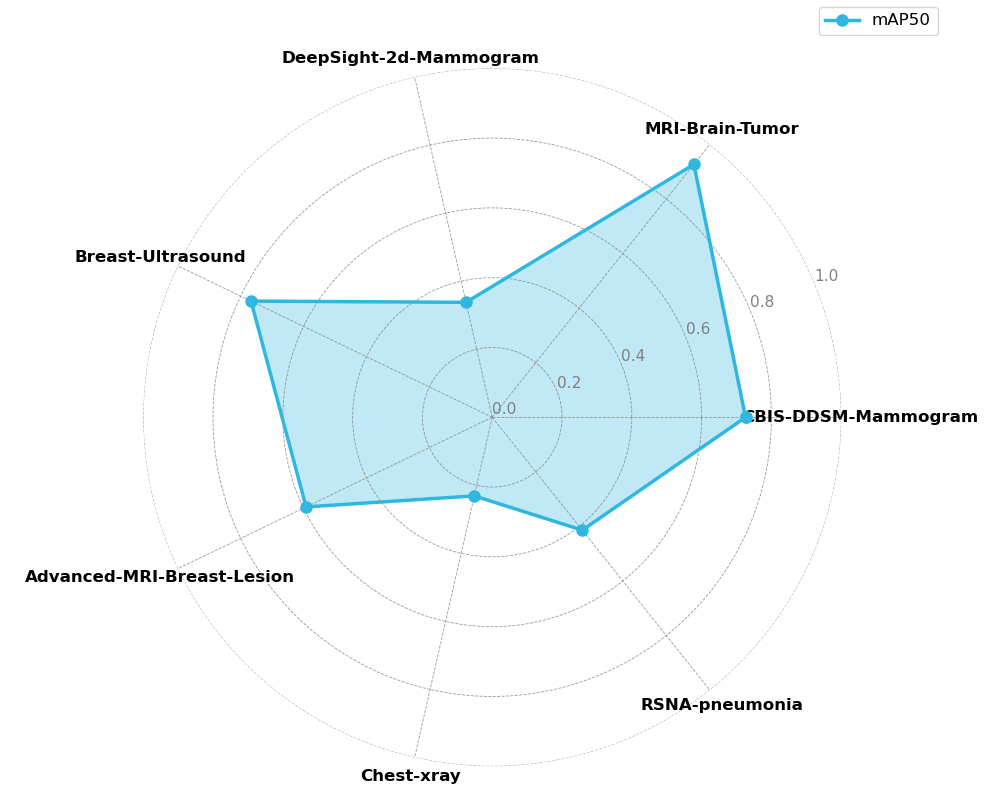
In contrast, the meta-learning approach successfully avoids catastrophic forgetting entirely, maintaining functional performance across all seven tasks simultaneously as shown in 10. This multi-task retention ability is particularly valuable in scenarios where process changes and domain variability introduce new tasks that must be incorporated into the training and deployment pipeline, while still preserving high performance on previously learned tasks
Experiment 6 - Few Shot Learning on novel EIT data
Table 4: Comparison of model performance between baseline transfer learning and meta-learning.
| Dataset | Baseline (%) | Meta-Model (%) | Δ (%) |
|---|---|---|---|
| mAP50 | mAP50 | ||
| EIT (n=30) | 62.9 | 87.5 | +24.6 |
| EIT (n=15) | 41.3 | 71.7 | +30.4 |
| Precision | Precision | ||
| EIT (n=30) | 64.6 | 86.1 | +21.5 |
| EIT (n=15) | 46.0 | 81.8 | +35.8 |
| Recall | Recall | ||
| EIT (n=30) | 61.4 | 90.1 | +28.7 |
| EIT (n=15) | 48.3 | 69.7 | +21.4 |
| F1-score | F1-score | ||
| EIT (n=30) | 62.3 | 88.9 | +26.6 |
| EIT (n=15) | 47.3 | 75.8 | +28.5 |
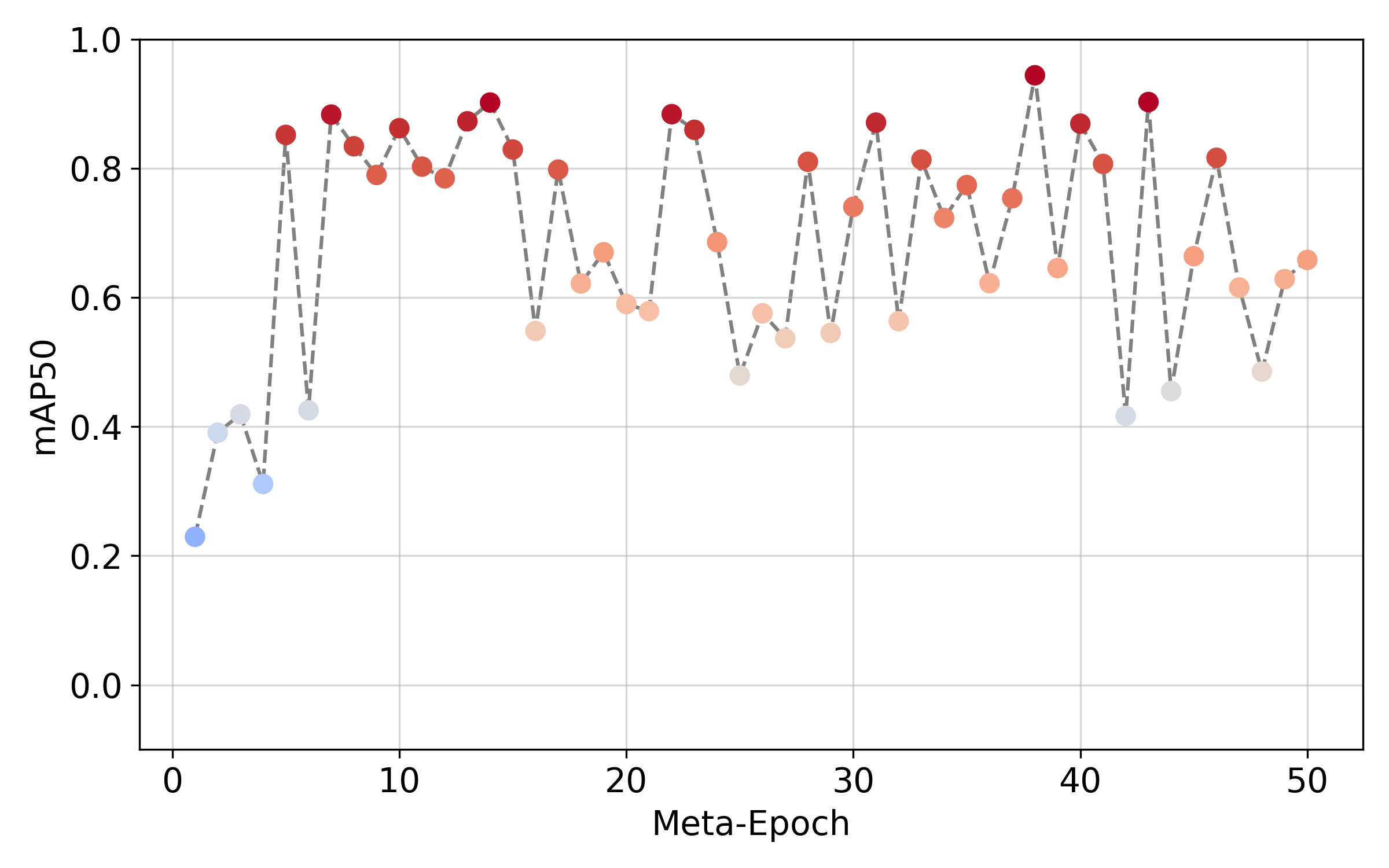
Few Shot Learning with n=30 samples
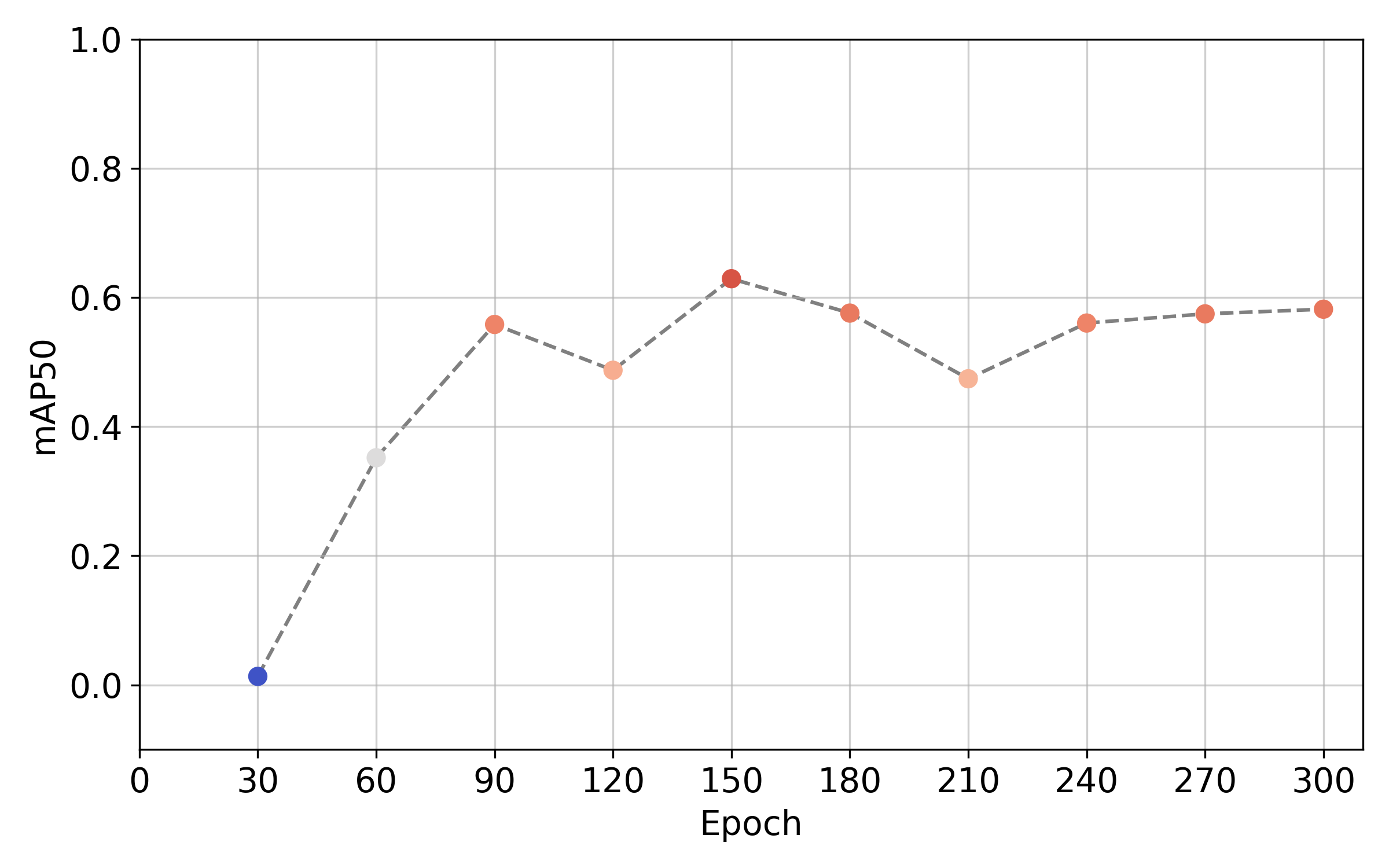
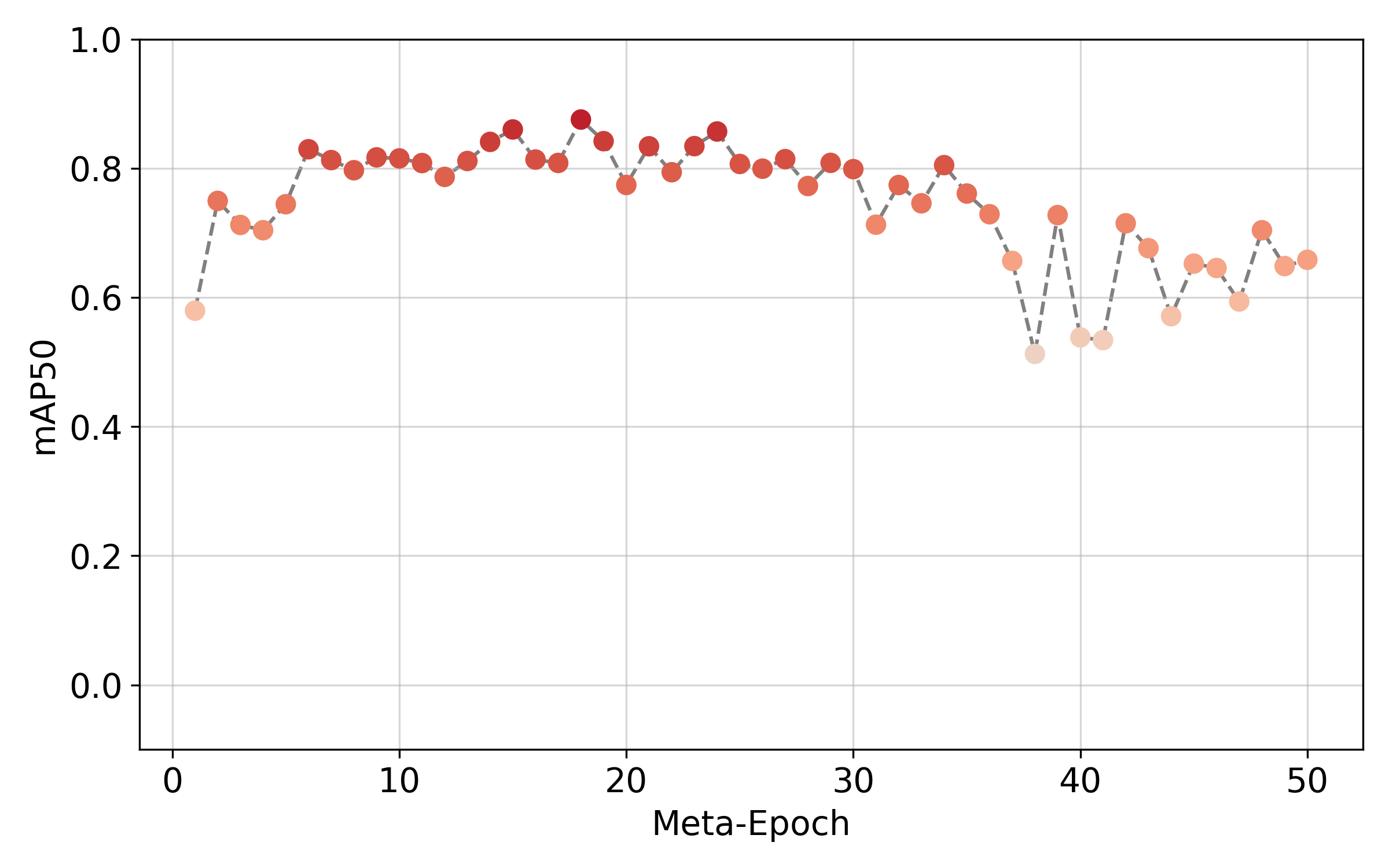
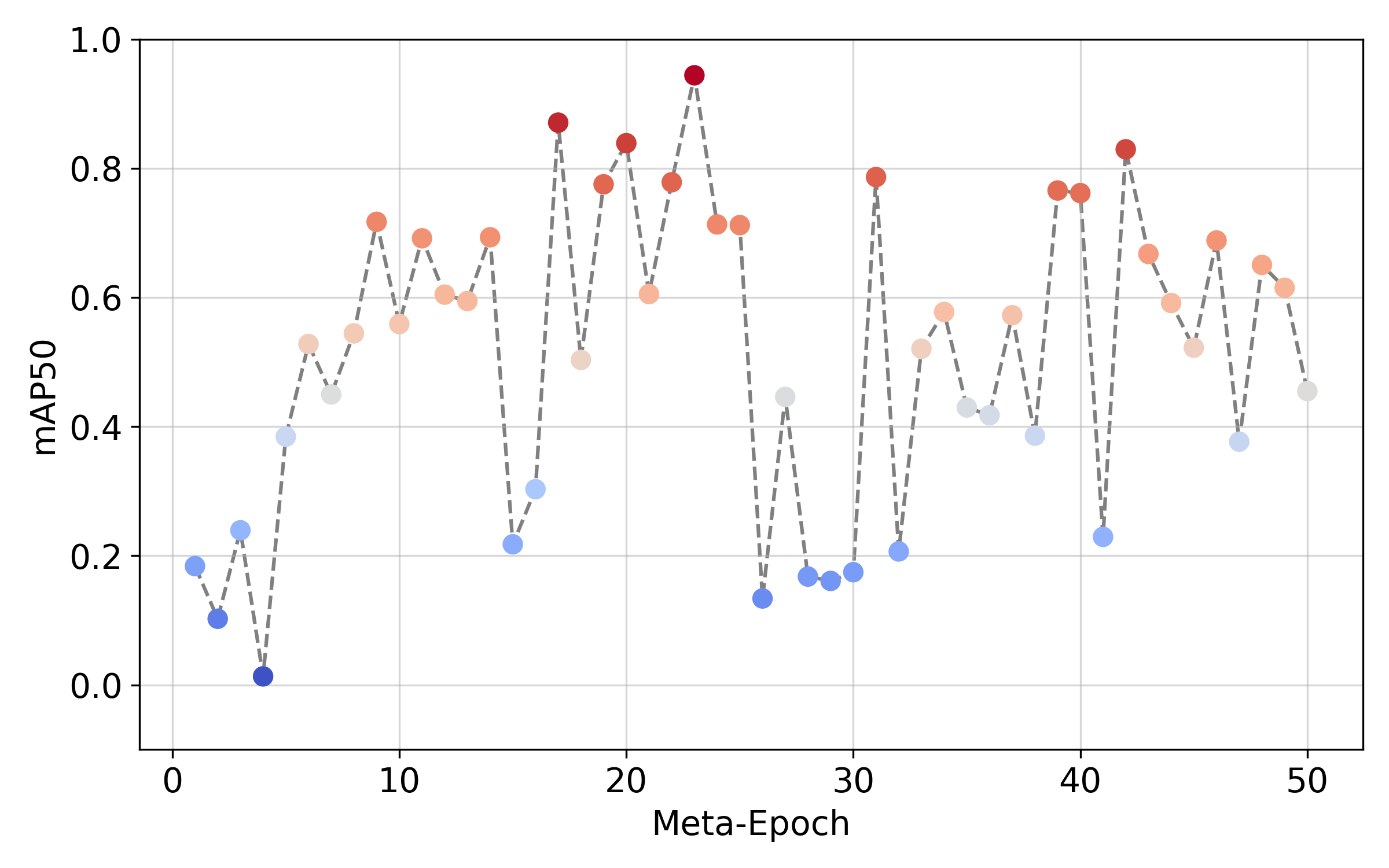
Very Few Shot Learning with n=15 samples
The meta-model obtained after training on 5 conventional tasks is finally fine-tuned on a very small EIT dataset consisting of 15 samples.
Conclusion
Systematic ablation studies yielded several key model insights. First, analysis of meta-learning rates established that a smaller rate (β=0.05) yielded the highest mAP50, balancing accuracy with robustness to noisy gradients. Second, experiments on the number and diversity of tasks used during training demonstrated that generalization follows an inverted U-shaped trend, with five tasks providing optimal meta-model performance, while additional out-of-distribution tasks led to negative transfer. Third, evaluation of base learner architectures revealed counterintuitive results, with smaller models outperforming larger ones trained without β-warmup, illustrating the importance of aligning network capacity with data scale to prevent overfitting. Fourth, experiments on fine-tuning schedules highlighted that, while fewer epochs could achieve >90% mAP50, they led to unstable performance, whereas extended fine-tuning improved consistency and reduced overfitting. Fifth, the meta-model achieved 88% mAP50 on limited EIT data versus 63% for the baseline model. Finally, compared to incremental transfer learning, the meta-model completely avoided catastrophic forgetting. Together, these results indicate that meta-learning can effectively leverage conventional imaging data to accelerate the development of models for novel modalities like EIT.
In conclusion, this research demonstrates that meta-learning, supported by a robust, extensible data processing pipeline, can effectively address the challenges of scarce training data in emerging medical imaging modalities such as EIT. Our framework enables very few-shot detection, leverages conventional imaging datasets to accelerate model development, and maintains robust performance across tasks without catastrophic forgetting. Systematic ablation studies highlight the importance of hyperparameter tuning, task selection and diversity, base learner network size and warmup schedules, and fine-tuning schedules for optimizing generalization. By combining open-source libraries for deterministic data handling and modular workflows, our approach provides a transparent, reproducible and extendable framework for future diagnostic applications. Collectively, these findings establish a generalizable blueprint for accelerating the clinical translation of novel imaging technologies and improving patient outcomes by leveraging meta-models that preserve knowledge across diverse diagnostic tasks, especially when collecting large datasets is impractical.
References
- R. L. Siegel, K. D. Miller, N. S. Wagle, and A. Jemal, “Cancer statistics, 2023,” CA Cancer J. Clin., vol. 73, no. 1, pp. 17–48, Jan. 2023, doi: 10.3322/CAAC.21763.
- L. A. Torre, F. Bray, R. L. Siegel, J. Ferlay, J. Lortet-Tieulent, and A. Jemal, “Global cancer statistics, 2012,” CA Cancer J. Clin., vol. 65, no. 2, pp. 87–108, Mar. 2015, doi: 10.3322/CAAC.21262.
- Milliman Research, “Lifetime health care costs for prevalent and preventable cancers,” The Mesothelioma Center, 2017. [Online]. Available: https://www.asbestos.com/featured-research/lifetime-healthcare-costs/
- C. Finn, P. Abbeel, and S. Levine, “Model-agnostic meta-learning for fast adaptation of deep networks,” in Proc. 34th Int. Conf. on Machine Learning (ICML’17), Sydney, NSW, Australia, 2017, pp. 1126–1135.
- A. Nichol, J. Achiam, and J. Schulman, “On First-Order Meta-Learning Algorithms,” ArXiv, vol. abs/1803.02999, 2018.
- T. Hospedales, A. Antoniou, P. Micaelli, and A. Storkey, “Meta-Learning in Neural Networks: A Survey,” IEEE Trans. Pattern Anal. Mach. Intell., vol. 44, no. 9, pp. 5149–5169, Sep. 2022, doi: 10.1109/TPAMI.2021.3079209.
- M. G. Marmot, D. G. Altman, D. A. Cameron, J. A. Dewar, S. G. Thompson, and M. Wilcox, “The benefits and harms of breast cancer screening: an independent review,” Br. J. Cancer, vol. 108, no. 11, pp. 2205–2240, Jun. 2013, doi: 10.1038/BJC.2013.177.
- G. K. Singh and A. Jemal, “Socioeconomic and racial/ethnic disparities in cancer mortality, incidence, and survival in the United States, 1950–2014: Over six decades of changing patterns and widening inequalities,” J. Environ. Public Health, vol. 2017, 2017, doi: 10.1155/2017/2819372.
- A. M. Myklebust, T. Seierstad, E. Stranden, and A. Lerdal, “Level of satisfaction during mammography screening in relation to discomfort, service provided, level of pain and breast compression,” Eur. J. Radiogr., vol. 1, no. 2, pp. 66–72, Jun. 2009.
- R. J. Halter, A. Hartov, and K. D. Paulsen, “A broadband high-frequency electrical impedance tomography system for breast imaging,” IEEE Trans. Biomed. Eng., vol. 55, no. 2 Pt 1, pp. 650–659, Feb. 2008, doi: 10.1109/TBME.2007.903516.
- B. Liu, B. Yang, C. Xu, J. Xia, M. Dai, Z. Ji, F. You, X. Dong, X. Shi, and F. Fu, “pyEIT: A Python-based framework for electrical impedance tomography,” SoftwareX, Oct. 2018, doi: 10.1016/j.softx.2018.07.002.
- G. Jocher, A. Chaurasia, and J. Qiu, “YOLOv8: Ultralytics Official Release,” Ultralytics, 2023.
- C. Villani, Optimal Transport: Old and New, Springer, 2009, doi: 10.1007/978-3-540-71050-9.
- L. van der Maaten and G. Hinton, “Visualizing data using t-SNE,” J. Mach. Learn. Res., vol. 9, pp. 2579–2605, Nov. 2008.


